dsDNA病毒中的蛋白质链甲变异。
IF 1
Q4 BIOPHYSICS
引用次数: 8
摘要
首先在噬菌体HK97中发现,生物链甲是由连接的蛋白质环形成的高度稳定的系统。环的每个亚基包含hk97样褶皱,其特征是其呈海底状,轴向(a)结构域为5链β片,外周(P)结构域为棘螺旋,延伸(E)环。HK97衣壳仅由一个HK97样折叠蛋白的共价连锁拷贝组成,代表了形成dsDNA基因组封装所需的高度稳定链甲的最有效策略。最近,通过低温电子显微镜(cryoEM)实现的近原子分辨率结构揭示了构建dsDNA病毒策略的一系列其他更复杂的变体。以p22样噬菌体为例,第一种策略是将插入(I)结构域附着在hk97样折叠的核心5链β片上。博德特拉噬菌体BPP-1的原子模型展示了HK97样折叠的经典HK97拓扑的另一种拓扑结构,以及构建稳定衣壳的第二种策略,其中辅助jellyroll蛋白二聚体用于巩固由衣壳蛋白亚基形成的非共价链甲。第三种策略是在类lambda噬菌体中发现的,它使用辅助蛋白三聚体来稳定3倍轴附近潜在的非共价链甲。疱疹病毒代表了高度复杂的病毒,它使用这些策略的组合,导致四层层次组织,包括由底区发现的hk97样折叠结构域形成的非共价链甲。对这些结构的透彻理解将有助于解开dsDNA病毒出现和进化的谜团,并为基于这些病毒的生物工程工作提供信息。本文章由计算机程序翻译,如有差异,请以英文原文为准。
Protein chainmail variants in dsDNA viruses.
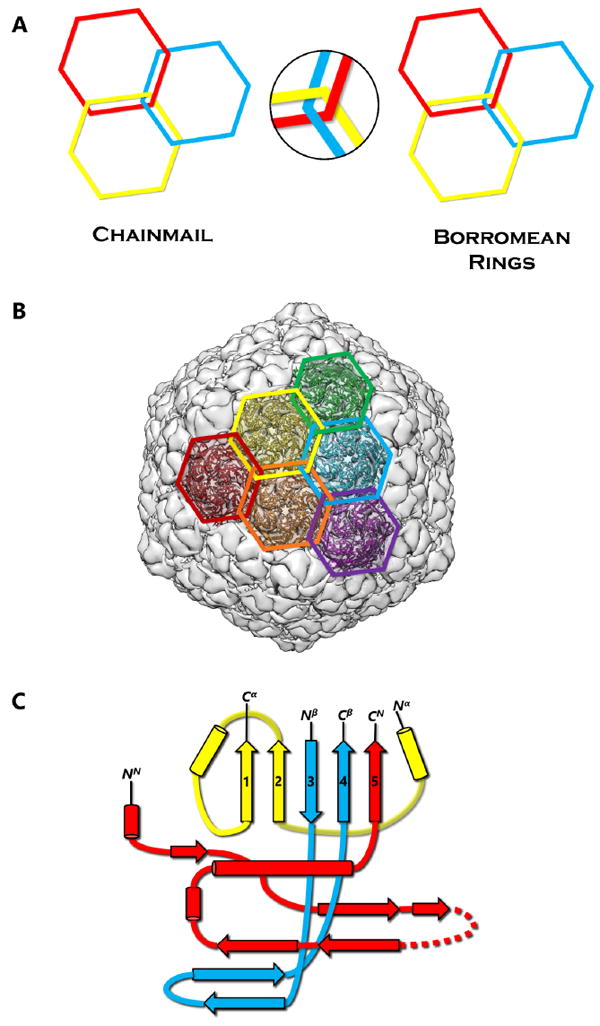
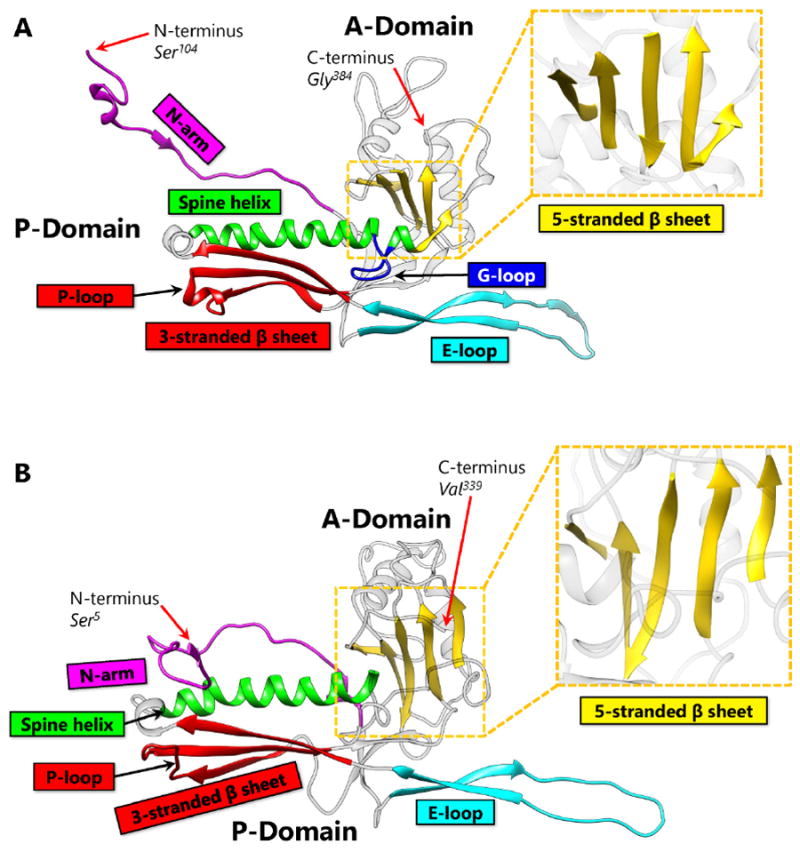
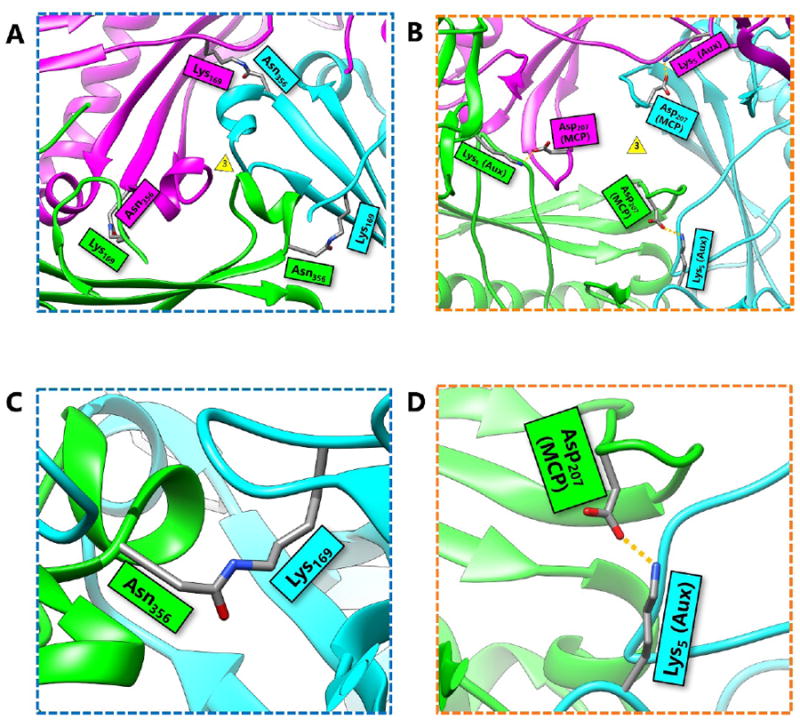
First discovered in bacteriophage HK97, biological chainmail is a highly stable system formed by concatenated protein rings. Each subunit of the ring contains the HK97-like fold, which is characterized by its submarine-like shape with a 5-stranded β sheet in the axial (A) domain, spine helix in the peripheral (P) domain, and an extended (E) loop. HK97 capsid consists of covalently-linked copies of just one HK97-like fold protein and represents the most effective strategy to form highly stable chainmail needed for dsDNA genome encapsidation. Recently, near-atomic resolution structures enabled by cryo electron microscopy (cryoEM) have revealed a range of other, more complex variants of this strategy for constructing dsDNA viruses. The first strategy, exemplified by P22-like phages, is the attachment of an insertional (I) domain to the core 5-stranded β sheet of the HK97-like fold. The atomic models of the Bordetella phage BPP-1 showcases an alternative topology of the classic HK97 topology of the HK97-like fold, as well as the second strategy for constructing stable capsids, where an auxiliary jellyroll protein dimer serves to cement the non-covalent chainmail formed by capsid protein subunits. The third strategy, found in lambda-like phages, uses auxiliary protein trimers to stabilize the underlying non-covalent chainmail near the 3-fold axis. Herpesviruses represent highly complex viruses that use a combination of these strategies, resulting in four-level hierarchical organization including a non-covalent chainmail formed by the HK97-like fold domain found in the floor region. A thorough understanding of these structures should help unlock the enigma of the emergence and evolution of dsDNA viruses and inform bioengineering efforts based on these viruses.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

AIMS Biophysics
BIOPHYSICS-
CiteScore
2.40
自引率
20.00%
发文量
16
审稿时长
8 weeks
期刊介绍:
AIMS Biophysics is an international Open Access journal devoted to publishing peer-reviewed, high quality, original papers in the field of biophysics. We publish the following article types: original research articles, reviews, editorials, letters, and conference reports. AIMS Biophysics welcomes, but not limited to, the papers from the following topics: · Structural biology · Biophysical technology · Bioenergetics · Membrane biophysics · Cellular Biophysics · Electrophysiology · Neuro-Biophysics · Biomechanics · Systems biology
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


