IMP2二聚化和RNA结合的结构见解
IF 2.7
3区 生物学
Q3 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
IGF2BP2 (IMP2)是一种与肿瘤发生和代谢紊乱有关的rna结合蛋白。针对单个IMP2结构域的结构研究为IMP2功能提供了重要的机制见解;然而,缺乏全长IMP2的结构信息,但这对于了解如何靶向药物发现中的IMP2活性是必要的。在这项研究中,我们利用生物物理和结构方法,包括质谱法、氢-氘交换耦合质谱法(HDX-MS)和小角度x射线散射法(SAXS),研究了全长IMP2的行为和RNA结合的影响。我们发现全长IMP2形成多种低聚态,但主要采用二聚体构象。从SAXS数据得出的分子模型表明,二聚体是由KH34和RRM1结构域以头到尾的方向形成的。在RNA结合后,IMP2形成与无载脂蛋白/RNA状态不同的伪对称二聚体。我们还发现,IMP2低聚物的形成,包括二聚体和高阶低聚物,对离子强度和RNA结合敏感。我们的发现首次深入了解了全长IMP2的结构特性,这可能为破坏其功能提供新的机会。本文章由计算机程序翻译,如有差异,请以英文原文为准。
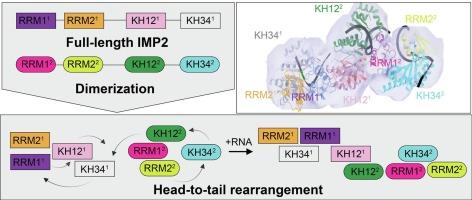
Structural insights into IMP2 dimerization and RNA binding
IGF2BP2 (IMP2) is an RNA-binding protein that contributes to tumorigenesis and metabolic disorders. Structural studies focused on individual IMP2 domains have provided important mechanistic insights into IMP2 function; however, structural information on full-length IMP2 is lacking but necessary to understand how to target IMP2 activity in drug discovery. In this study, we investigated the behavior of full-length IMP2 and the influence of RNA binding using biophysical and structural methods including mass photometry, hydrogen–deuterium exchange coupled to mass spectrometry (HDX-MS), and small angle x-ray scattering (SAXS). We found that full-length IMP2 forms multiple oligomeric states but predominantly adopts a dimeric conformation. Molecular models derived from SAXS data suggest the dimer is formed in a head-to-tail orientation by the KH34 and RRM1 domains. Upon RNA binding, IMP2 forms a pseudo-symmetric dimer different from its apo/RNA-free state. We also found that the formation of IMP2 oligomeric species, which includes dimers and higher-order oligomers, is sensitive to ionic strength and RNA binding. Our findings provide the first insight into the structural properties of full-length IMP2, which may lead to novel opportunities for disrupting its function.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Journal of structural biology
生物-生化与分子生物学
CiteScore
6.30
自引率
3.30%
发文量
88
审稿时长
65 days
期刊介绍:
Journal of Structural Biology (JSB) has an open access mirror journal, the Journal of Structural Biology: X (JSBX), sharing the same aims and scope, editorial team, submission system and rigorous peer review. Since both journals share the same editorial system, you may submit your manuscript via either journal homepage. You will be prompted during submission (and revision) to choose in which to publish your article. The editors and reviewers are not aware of the choice you made until the article has been published online. JSB and JSBX publish papers dealing with the structural analysis of living material at every level of organization by all methods that lead to an understanding of biological function in terms of molecular and supermolecular structure.
Techniques covered include:
• Light microscopy including confocal microscopy
• All types of electron microscopy
• X-ray diffraction
• Nuclear magnetic resonance
• Scanning force microscopy, scanning probe microscopy, and tunneling microscopy
• Digital image processing
• Computational insights into structure
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


