IF 14.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
由小型开放阅读框编码的微蛋白是蛋白质组中的 "暗物质"。尽管在生命三大领域的各种生物体中都检测到了微蛋白,但仍有许多微蛋白有待鉴定,而且只有少数微蛋白具有功能特征。在这项关于肠杆菌科 5,668 个细菌基因组中基因间小开放阅读框(ismORFs,15-70 个密码子)的综合研究中,我们发现了 67,297 个经过纯化选择的 ismORFs 簇。在测试的 16 个ismORFs 中,有 11 个检测到了标记大肠杆菌微蛋白的表达,验证了预测结果。虽然 ismORFs 主要编码疏水性、潜在跨膜、非结构化或最小结构化的微蛋白,但也预测了一些球状褶皱、寡聚体结构以及可能与邻近基因编码的蛋白发生的相互作用。关于预测的微蛋白家族的完整信息,包括转录和翻译的证据以及结构预测,可作为研究微蛋白功能的简易搜索资源。本文章由计算机程序翻译,如有差异,请以英文原文为准。
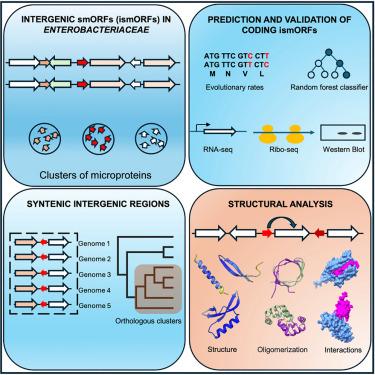
The hidden bacterial microproteome
Microproteins encoded by small open reading frames comprise the “dark matter” of proteomes. Although microproteins have been detected in diverse organisms from all three domains of life, many more remain to be identified, and only a few have been functionally characterized. In this comprehensive study of intergenic small open reading frames (ismORFs, 15–70 codons) in 5,668 bacterial genomes of the family Enterobacteriaceae, we identify 67,297 clusters of ismORFs subject to purifying selection. Expression of tagged Escherichia coli microproteins is detected for 11 of the 16 tested, validating the predictions. Although the ismORFs mainly code for hydrophobic, potentially transmembrane, unstructured, or minimally structured microproteins, some globular folds, oligomeric structures, and possible interactions with proteins encoded by neighboring genes are predicted. Complete information on the predicted microprotein families, including evidence of transcription and translation, and structure predictions are available as an easily searchable resource for investigation of microprotein functions.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


