单分子m6A检测通过内源性标记揭示了RNA异构体的复杂性
IF 14.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
n6 -甲基腺苷(m6A)在不同RNA异构体上的分布尚不完全清楚。在HEK293T细胞中,我们通过apobec1 - yth诱导的C-to-U突变,在10-100 nt外的单牛津纳米孔技术(ONT)直接RNA测序(DRS)读取上内源性标记甲基化的m6A位点,获得1,020,237个5-mer单读m6A信号。然后,我们训练了m6Aiso,这是一种深度残差神经网络模型,可以在单读分辨率下准确识别和量化m6A。分析m6aiso测定的m6A单读和同工异构体,揭示了m6A位点沿单个分子的距离依赖键。它还揭示了内含子保留异构体上相同的m6A位点的特异性甲基化,部分原因是它们与外显子连接处的距离不同以及与TARBP2的异构体特异性结合。此外,我们发现转录因子SMAD3在上皮-间质转化过程中促进m6A在其转录RNA亚型上的沉积,从而导致m6A在具有替代启动子的亚型上的特异性调控。我们的研究强调了m6Aiso在阐明m6A跨RNA异构体的复杂动力学和复杂性方面的有效性。本文章由计算机程序翻译,如有差异,请以英文原文为准。
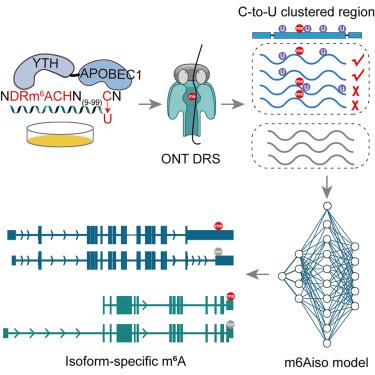
Single-molecule m6A detection empowered by endogenous labeling unveils complexities across RNA isoforms
The landscape of N6-methyadenosine (m6A) on different RNA isoforms is still incompletely understood. Here, in HEK293T cells, we endogenously label the methylated m6A sites on single Oxford Nanopore Technology (ONT) direct RNA sequencing (DRS) reads by APOBEC1-YTH-induced C-to-U mutations 10–100 nt away, obtaining 1,020,237 5-mer single-read m6A signals. We then trained m6Aiso, a deep residual neural network model that accurately identifies and quantifies m6A at single-read resolution. Analyzing m6Aiso-determined m6A on single reads and isoforms uncovers distance-dependent linkages of m6A sites along single molecules. It also uncovers specific methylation of identical m6A sites on intron-retained isoforms, partly due to their differential distances to exon junctions and isoform-specific binding of TARBP2. Moreover, we find that transcription factor SMAD3 promotes m6A deposition on its transcribed RNA isoforms during epithelial-mesenchymal transition, resulting in isoform-specific regulation of m6A on isoforms with alternative promoters. Our study underscores the effectiveness of m6Aiso in elucidating the intricate dynamics and complexities of m6A across RNA isoforms.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


