表型驱动的分子遗传测试建议用于诊断儿科罕见疾病
IF 12.4
1区 医学
Q1 HEALTH CARE SCIENCES & SERVICES
引用次数: 0
摘要
罕见病患者往往会经历长时间的诊断延误。进行适当的基因检测至关重要,但也极具挑战性,尤其是对于没有遗传学专业知识的普通儿科医生而言。美国医学遗传学会(ACMG)最近的指南支持尽早使用外显子组测序(ES)或基因组测序(GS)来诊断先天性异常或发育迟缓等疾病,但同时仍建议对表现出特定疾病强烈表现的患者进行基因检测。我们认识到浏览这些选项的难度,因此开发了一个机器学习模型,该模型以哥伦比亚大学欧文医学中心的 1005 份患者记录为训练对象,根据表型信息推荐适当的基因测试。该模型的AUROC为0.823,AUPRC为0.918,与遗传学专家的决定非常一致,表现出了卓越的性能,并在外部队列中表现出了很强的普适性(AUROC:0.77,AUPRC:0.816),这表明它对普通儿科医生来说具有潜在价值,可以通过加强基因测试订购来加快罕见病的诊断。本文章由计算机程序翻译,如有差异,请以英文原文为准。
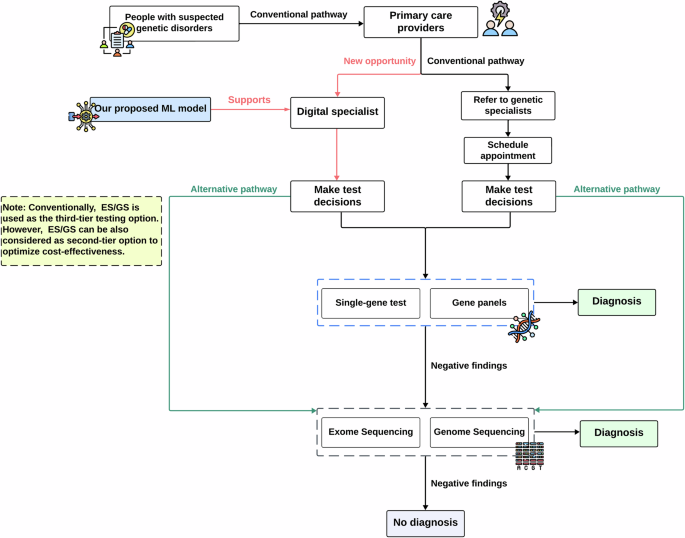
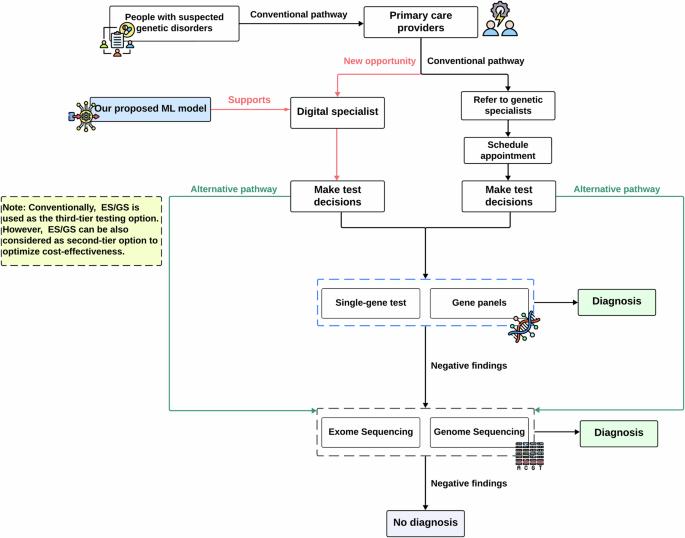
Phenotype driven molecular genetic test recommendation for diagnosing pediatric rare disorders
Patients with rare diseases often experience prolonged diagnostic delays. Ordering appropriate genetic tests is crucial yet challenging, especially for general pediatricians without genetic expertise. Recent American College of Medical Genetics (ACMG) guidelines embrace early use of exome sequencing (ES) or genome sequencing (GS) for conditions like congenital anomalies or developmental delays while still recommend gene panels for patients exhibiting strong manifestations of a specific disease. Recognizing the difficulty in navigating these options, we developed a machine learning model trained on 1005 patient records from Columbia University Irving Medical Center to recommend appropriate genetic tests based on the phenotype information. The model achieved a remarkable performance with an AUROC of 0.823 and AUPRC of 0.918, aligning closely with decisions made by genetic specialists, and demonstrated strong generalizability (AUROC:0.77, AUPRC: 0.816) in an external cohort, indicating its potential value for general pediatricians to expedite rare disease diagnosis by enhancing genetic test ordering.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

NPJ Digital Medicine
Multiple-
CiteScore
25.10
自引率
3.30%
发文量
170
审稿时长
15 weeks
期刊介绍:
npj Digital Medicine is an online open-access journal that focuses on publishing peer-reviewed research in the field of digital medicine. The journal covers various aspects of digital medicine, including the application and implementation of digital and mobile technologies in clinical settings, virtual healthcare, and the use of artificial intelligence and informatics.
The primary goal of the journal is to support innovation and the advancement of healthcare through the integration of new digital and mobile technologies. When determining if a manuscript is suitable for publication, the journal considers four important criteria: novelty, clinical relevance, scientific rigor, and digital innovation.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


