2024 年的 BioPAX:我们的现状和未来
IF 4.1
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
Computational and structural biotechnology journal
Pub Date : 2024-11-04
DOI:10.1016/j.csbj.2024.10.045
引用次数: 0
摘要
在系统生物学中,生物通路研究在了解生物系统的复杂性方面发挥着核心作用。近年来,众多在线数据库提供了大量的通路数据,因此亟需对这些数据进行标准化处理。BioPAX 格式(生物通路交换)于 2010 年出现,是跨数据库标准化和交换通路数据的解决方案。BioPAX 是一种与本体相关联的语义网格式。它具有很强的表达能力,可以在分子和细胞层面精细地描述生物通路,但相关的内在复杂性可能会成为其被广泛采用的障碍。在此,我们报告了 BioPAX 格式在 2024 年的使用情况。在此,我们将报告 2024 年 BioPAX 格式的使用情况。我们将比较不同的通路数据库如何使用 BioPAX 来规范其数据,并指出可能的改进途径,以充分利用其潜力。我们还报告了为处理 BioPAX 数据而开发的各种工具和软件。最后,我们提出了一种新的 BioPAX 图抽象概念,它可以专门针对特定分析所需的 BioPAX 图区域,从而区分适合表示的格式和适合上下文分析的抽象概念。本文章由计算机程序翻译,如有差异,请以英文原文为准。
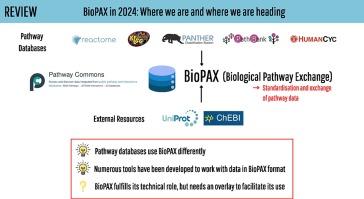
BioPAX in 2024: Where we are and where we are heading
In systems biology, the study of biological pathways plays a central role in understanding the complexity of biological systems. The massification of pathway data made available by numerous online databases in recent years has given rise to an important need for standardization of this data. The BioPAX format (Biological Pathway Exchange) emerged in 2010 as a solution for standardizing and exchanging pathway data across databases. BioPAX is a Semantic Web format associated to an ontology. It is highly expressive, allowing to finely describe biological pathways at the molecular and cellular levels, but the associated intrinsic complexity may be an obstacle to its widespread adoption.
Here, we report on the use of the BioPAX format in 2024. We compare how the different pathway databases use BioPAX to standardize their data and point out possible avenues for improvement to make full use of its potential. We also report on the various tools and software that have been developed to work with BioPAX data. Finally, we present a new concept of abstraction on BioPAX graphs that would allow to specifically target areas in a BioPAX graph needed for a specific analysis, thus differentiating the format suited for representation and the abstraction suited for contextual analysis.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Computational and structural biotechnology journal
Biochemistry, Genetics and Molecular Biology-Biophysics
CiteScore
9.30
自引率
3.30%
发文量
540
审稿时长
6 weeks
期刊介绍:
Computational and Structural Biotechnology Journal (CSBJ) is an online gold open access journal publishing research articles and reviews after full peer review. All articles are published, without barriers to access, immediately upon acceptance. The journal places a strong emphasis on functional and mechanistic understanding of how molecular components in a biological process work together through the application of computational methods. Structural data may provide such insights, but they are not a pre-requisite for publication in the journal. Specific areas of interest include, but are not limited to:
Structure and function of proteins, nucleic acids and other macromolecules
Structure and function of multi-component complexes
Protein folding, processing and degradation
Enzymology
Computational and structural studies of plant systems
Microbial Informatics
Genomics
Proteomics
Metabolomics
Algorithms and Hypothesis in Bioinformatics
Mathematical and Theoretical Biology
Computational Chemistry and Drug Discovery
Microscopy and Molecular Imaging
Nanotechnology
Systems and Synthetic Biology
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


