微处理器与变体 NEXT 复合物之间的功能连接
IF 14.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
在哺乳动物细胞中,初级miRNA在其发夹结构处被Microprocessor复合体裂解,该复合体的核心由DROSHA和DGCR8组成。在这里,我们发现,在小鼠胚胎干细胞(mESCs)中,由微处理器裂解产生的5′侧翼区域是RNA外泌体的靶标。DGCR8与核外泌体靶向(NEXT)元件ZCCHC8之间的物理联系促进了这一过程。然而,令人惊讶的是,生化研究和诱变研究都表明,含有 RNA 螺旋酶 MTR4 但没有 RNA 结合蛋白 RBM7 的变体 NEXT 复合物才是活跃的实体。这种微处理器-NEXT变体也以其他基因组区域(如增强子)表达的含茎环 RNA 为靶标。相比之下,Microprocessor 不参与结构较少的 NEXT 底物的周转。因此,我们的研究结果表明,MTR4-ZCCHC8 可以与 RBM7 或 DGCR8/DROSHA 连接,根据不同的结构背景靶向不同的 RNA 底物。本文章由计算机程序翻译,如有差异,请以英文原文为准。
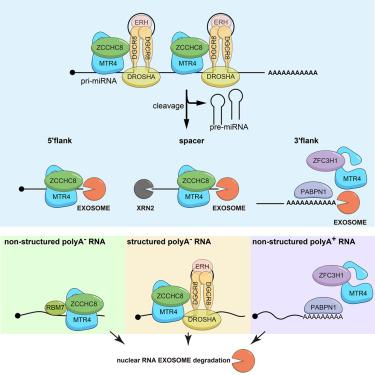
A functional connection between the Microprocessor and a variant NEXT complex
In mammalian cells, primary miRNAs are cleaved at their hairpin structures by the Microprocessor complex, whose core is composed of DROSHA and DGCR8. Here, we show that 5′ flanking regions, resulting from Microprocessor cleavage, are targeted by the RNA exosome in mouse embryonic stem cells (mESCs). This is facilitated by a physical link between DGCR8 and the nuclear exosome targeting (NEXT) component ZCCHC8. Surprisingly, however, both biochemical and mutagenesis studies demonstrate that a variant NEXT complex, containing the RNA helicase MTR4 but devoid of the RNA-binding protein RBM7, is the active entity. This Microprocessor-NEXT variant also targets stem-loop-containing RNAs expressed from other genomic regions, such as enhancers. By contrast, Microprocessor does not contribute to the turnover of less structured NEXT substrates. Our results therefore demonstrate that MTR4-ZCCHC8 can link to either RBM7 or DGCR8/DROSHA to target different RNA substrates depending on their structural context.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


