GEMimp:利用图嵌入神经网络对微生物组数据进行准确而稳健的估算方法。
IF 4.7
2区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
微生物组研究日益凸显微生物组成与人类健康之间的深刻联系,大量研究证实微生物组特征与各种疾病之间存在密切联系。然而,微生物组数据的分析经常受到固有稀疏性问题的影响,其特点是存在大量观测到的零。这些零不仅扭曲了微生物物种的丰度分布,还破坏了从这些数据中得出的科学结论的可靠性。为了应对这一挑战,我们引入了 GEMimp,这是一种创新的估算方法,旨在为微生物组数据分析注入稳健性。GEMimp 利用了 node2vec 算法,该算法在随机游走采样过程中同时采用了广度优先搜索(BFS)和深度优先搜索(DFS)策略。这种方法使 GEMimp 能够学习每个分类单元的细微、低维表征,从而以前所未有的准确性重建它们的相似性网络。我们将 GEMimp 与最先进的估算方法(包括 SAVER、MAGIC 和 mbImpute)进行了比较分析。结果清楚地表明,与原始数据集相比,GEMimp 取得了最高的皮尔逊相关系数,表现优于同类方法。此外,GEMimp 在识别重要类群、增强疾病相关类群的检测以及有效减轻稀疏性对模拟数据集和真实数据集(如与 2 型糖尿病(T2D)和结直肠癌(CRC)相关的数据集)的影响方面表现出了显著的能力。这些发现共同凸显了 GEMimp 的强大功效,使其能够更好地分析微生物数据。随着稀疏性问题的缓解,它可以极大地促进下游分析,甚至是微生物学领域的分析。本文章由计算机程序翻译,如有差异,请以英文原文为准。
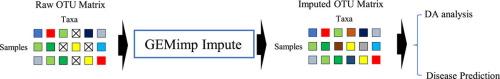
GEMimp: An Accurate and Robust Imputation Method for Microbiome Data Using Graph Embedding Neural Network
Microbiome research has increasingly underscored the profound link between microbial compositions and human health, with numerous studies establishing a strong correlation between microbiome characteristics and various diseases. However, the analysis of microbiome data is frequently compromised by inherent sparsity issues, characterized by a substantial presence of observed zeros. These zeros not only skew the abundance distribution of microbial species but also undermine the reliability of scientific conclusions drawn from such data. Addressing this challenge, we introduce GEMimp, an innovative imputation method designed to infuse robustness into microbiome data analysis. GEMimp leverages the node2vec algorithm, which incorporates both Breadth-First Search (BFS) and Depth-First Search (DFS) strategies in its random walks sampling process. This approach enables GEMimp to learn nuanced, low-dimensional representations of each taxonomic unit, facilitating the reconstruction of their similarity networks with unprecedented accuracy.
Our comparative analysis pits GEMimp against state-of-the-art imputation methods including SAVER, MAGIC and mbImpute. The results unequivocally demonstrate that GEMimp outperforms its counterparts by achieving the highest Pearson correlation coefficient when compared to the original raw dataset. Furthermore, GEMimp shows notable proficiency in identifying significant taxa, enhancing the detection of disease-related taxa and effectively mitigating the impact of sparsity on both simulated and real-world datasets, such as those pertaining to Type 2 Diabetes (T2D) and Colorectal Cancer (CRC). These findings collectively highlight the strong effectiveness of GEMimp, allowing for better analysis on microbial data. With alleviation of sparsity issues, it could be greatly facilitated in downstream analyses and even in the field of microbiology.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
Journal of Molecular Biology
生物-生化与分子生物学
CiteScore
11.30
自引率
1.80%
发文量
412
审稿时长
28 days
期刊介绍:
Journal of Molecular Biology (JMB) provides high quality, comprehensive and broad coverage in all areas of molecular biology. The journal publishes original scientific research papers that provide mechanistic and functional insights and report a significant advance to the field. The journal encourages the submission of multidisciplinary studies that use complementary experimental and computational approaches to address challenging biological questions.
Research areas include but are not limited to: Biomolecular interactions, signaling networks, systems biology; Cell cycle, cell growth, cell differentiation; Cell death, autophagy; Cell signaling and regulation; Chemical biology; Computational biology, in combination with experimental studies; DNA replication, repair, and recombination; Development, regenerative biology, mechanistic and functional studies of stem cells; Epigenetics, chromatin structure and function; Gene expression; Membrane processes, cell surface proteins and cell-cell interactions; Methodological advances, both experimental and theoretical, including databases; Microbiology, virology, and interactions with the host or environment; Microbiota mechanistic and functional studies; Nuclear organization; Post-translational modifications, proteomics; Processing and function of biologically important macromolecules and complexes; Molecular basis of disease; RNA processing, structure and functions of non-coding RNAs, transcription; Sorting, spatiotemporal organization, trafficking; Structural biology; Synthetic biology; Translation, protein folding, chaperones, protein degradation and quality control.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


