通过网络沙普利值分析发现乳腺癌抗 PD1 反应的可解释生物标志物
IF 4.9
2区 医学
Q1 COMPUTER SCIENCE, INTERDISCIPLINARY APPLICATIONS
引用次数: 0
摘要
背景和目的:免疫疗法有望提高乳腺癌的病理完全反应率,但仅限于部分患者。因此,找出预测治疗反应性的因素至关重要。基因表达和调控本身就在错综复杂的网络中运行,构成了细胞过程的基本分子机制,通常可作为可靠的生物标志物。方法:为了解决这些局限性,我们设计了一种基于合作博弈论的新型特征选择方法,并与复杂的机器学习模型进行了和谐整合。这种方法能识别具有先验遗传关联结构的相互关联的基因调控网络生物标记模块。具体来说,我们利用网络上的夏普利(Shapley)值来量化特征的重要性,同时根据网络扩展原则和节点邻接关系对其整合进行策略性限制,从而提高特征选择的可解释性。我们将我们的方法应用于乳腺癌免疫疗法反应的公开单细胞 RNA 测序数据集,并将确定的特征基因集作为生物标记物。独立验证的功能富集分析进一步说明了这些方法的有效预测性能 结果:我们证明了所提出的方法在具有网络结构的数据中的复杂性和卓越性。它揭示了一个包含 27 个免疫疗法反应基因的内聚生物标志物模块。值得注意的是,该模块能够精确预测乳腺癌患者的抗 PD1 治疗结果,其分类准确率为 0.905,AUC 值为 0.971,凸显了其揭示基因功能的独特能力 结论:所提出的方法能有效识别网络模块生物标记物,检测到的抗 PD1 反应生物标记物能丰富我们对免疫疗法潜在生理机制的理解,在实现精准医疗方面具有广阔的应用前景。本文章由计算机程序翻译,如有差异,请以英文原文为准。
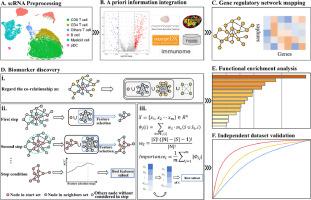
Discovering explainable biomarkers for breast cancer anti-PD1 response via network Shapley value analysis
Background and objective
Immunotherapy holds promise in enhancing pathological complete response rates in breast cancer, albeit confined to a select cohort of patients. Consequently, pinpointing factors predictive of treatment responsiveness is of paramount importance. Gene expression and regulation, inherently operating within intricate networks, constitute fundamental molecular machinery for cellular processes and often serve as robust biomarkers. Nevertheless, contemporary feature selection approaches grapple with two key challenges: opacity in modeling and scarcity in accounting for gene-gene interactions
Methods
To address these limitations, we devise a novel feature selection methodology grounded in cooperative game theory, harmoniously integrating with sophisticated machine learning models. This approach identifies interconnected gene regulatory network biomarker modules with priori genetic linkage architecture. Specifically, we leverage Shapley values on network to quantify feature importance, while strategically constraining their integration based on network expansion principles and nodal adjacency, thereby fostering enhanced interpretability in feature selection. We apply our methods to a publicly available single-cell RNA sequencing dataset of breast cancer immunotherapy responses, using the identified feature gene set as biomarkers. Functional enrichment analysis with independent validations further illustrates their effective predictive performance
Results
We demonstrate the sophistication and excellence of the proposed method in data with network structure. It unveiled a cohesive biomarker module encompassing 27 genes for immunotherapy response. Notably, this module proves adept at precisely predicting anti-PD1 therapeutic outcomes in breast cancer patients with classification accuracy of 0.905 and AUC value of 0.971, underscoring its unique capacity to illuminate gene functionalities
Conclusion
The proposed method is effective for identifying network module biomarkers, and the detected anti-PD1 response biomarkers can enrich our understanding of the underlying physiological mechanisms of immunotherapy, which have a promising application for realizing precision medicine.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Computer methods and programs in biomedicine
工程技术-工程:生物医学
CiteScore
12.30
自引率
6.60%
发文量
601
审稿时长
135 days
期刊介绍:
To encourage the development of formal computing methods, and their application in biomedical research and medical practice, by illustration of fundamental principles in biomedical informatics research; to stimulate basic research into application software design; to report the state of research of biomedical information processing projects; to report new computer methodologies applied in biomedical areas; the eventual distribution of demonstrable software to avoid duplication of effort; to provide a forum for discussion and improvement of existing software; to optimize contact between national organizations and regional user groups by promoting an international exchange of information on formal methods, standards and software in biomedicine.
Computer Methods and Programs in Biomedicine covers computing methodology and software systems derived from computing science for implementation in all aspects of biomedical research and medical practice. It is designed to serve: biochemists; biologists; geneticists; immunologists; neuroscientists; pharmacologists; toxicologists; clinicians; epidemiologists; psychiatrists; psychologists; cardiologists; chemists; (radio)physicists; computer scientists; programmers and systems analysts; biomedical, clinical, electrical and other engineers; teachers of medical informatics and users of educational software.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


