mTORC1 通过自动调节 TFE3 在溶酶体上的存在来限制其活性
IF 16.6
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
摘要
要刺激细胞生长,蛋白激酶复合物 mTORC1 需要细胞内氨基酸才能激活。氨基酸充足时,溶酶体上的 Rag GTP 酶会向 mTORC1 转运氨基酸,生长因子信号通过 GTP 酶 Rheb 增强 mTORC1 的活性。在缺乏氨基酸的情况下,GATOR1 会使 Rag 失活,导致溶酶体脱离和 mTORC1 失活。我们证明,在人体细胞中,mTORC1 从溶酶体的释放取决于其激酶活性。根据负反馈机制,激活的 mTOR 突变体显示出较低的溶酶体占有率,导致溶酶体底物 TFE3 的低磷酸化和核定位。令人惊讶的是,被 Rheb 激活的 mTORC1 并不增加 mTORC1 的细胞质/溶酶体比例,这表明存在具有不同底物特异性的 mTORC1 池。任何一个池的失调都会导致 TFE3 活性异常,这可能是局灶性皮质发育不良 II 型(FCD II)和结节性硬化症(TSC)致痫畸形中 TFE3 核聚集的原因。本文章由计算机程序翻译,如有差异,请以英文原文为准。
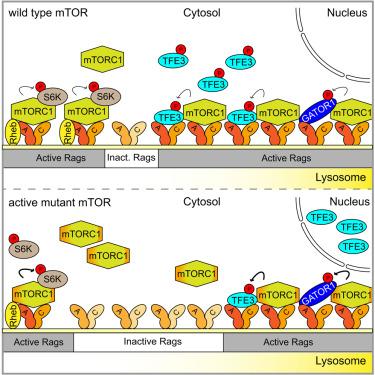
mTORC1 restricts TFE3 activity by auto-regulating its presence on lysosomes
To stimulate cell growth, the protein kinase complex mTORC1 requires intracellular amino acids for activation. Amino-acid sufficiency is relayed to mTORC1 by Rag GTPases on lysosomes, where growth factor signaling enhances mTORC1 activity via the GTPase Rheb. In the absence of amino acids, GATOR1 inactivates the Rags, resulting in lysosomal detachment and inactivation of mTORC1. We demonstrate that in human cells, the release of mTORC1 from lysosomes depends on its kinase activity. In accordance with a negative feedback mechanism, activated mTOR mutants display low lysosome occupancy, causing hypo-phosphorylation and nuclear localization of the lysosomal substrate TFE3. Surprisingly, mTORC1 activated by Rheb does not increase the cytoplasmic/lysosomal ratio of mTORC1, indicating the existence of mTORC1 pools with distinct substrate specificity. Dysregulation of either pool results in aberrant TFE3 activity and may explain nuclear accumulation of TFE3 in epileptogenic malformations in focal cortical dysplasia type II (FCD II) and tuberous sclerosis (TSC).
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


