利用深度学习模型改进扫描实验室报告中的表格数据提取。
IF 4
2区 医学
Q2 COMPUTER SCIENCE, INTERDISCIPLINARY APPLICATIONS
引用次数: 0
摘要
目的:医学实验室检测在医疗保健中至关重要,可为诊断和治疗提供关键数据。然而,病人的化验结果通常是通过传真在各医疗机构之间传递的,无法立即用于及时的临床决策。因此,开发新技术从扫描的化验报告中准确提取化验信息非常重要。本研究旨在开发一种基于深度学习的先进光学字符识别(OCR)方法,以识别扫描化验报告中包含化验结果的表格:从扫描的实验报告中提取表格数据涉及两个阶段:表格检测(即识别表格对象的区域)和表格识别(即识别和提取表格结构和内容)。DETR R18 算法和 YOLOv8s 参与了表格检测,我们比较了 PaddleOCR 和编码器-双解码器(EDD)模型在表格识别方面的性能。我们对随机抽取的 632 份实验室测试报告中的 650 张桌子进行了标注,并用于训练和评估这些模型。在表格检测评估中,我们使用了平均精度 (AP)、平均召回率 (AR)、AP50 和 AP75 等指标。在表格识别评估中,我们使用了树-编辑距离(Tree-Edit Distance,TEDS):结果:在表格检测方面,微调后的 DETR R18 表现优异(AP50:0.774;AP75:0.644;AP:0.601;AR:0.766)。在表格识别方面,微调 EDD 的 TEDS 得分为 0.815,优于其他模型。拟议的 OCR 管道(微调 DETR R18 和微调 EDD)取得了令人印象深刻的结果,TEDS 得分为 0.699,TEDS 结构得分为 0.764:我们的研究为扫描的临床文件提供了一个专用的 OCR 管道,利用最先进的深度学习模型进行兴趣区域检测和表格识别。高 TEDS 分数证明了我们方法的有效性,这对临床数据分析和决策具有重要意义。本文章由计算机程序翻译,如有差异,请以英文原文为准。
Improving tabular data extraction in scanned laboratory reports using deep learning models
Objective
Medical laboratory testing is essential in healthcare, providing crucial data for diagnosis and treatment. Nevertheless, patients’ lab testing results are often transferred via fax across healthcare organizations and are not immediately available for timely clinical decision making. Thus, it is important to develop new technologies to accurately extract lab testing information from scanned laboratory reports. This study aims to develop an advanced deep learning-based Optical Character Recognition (OCR) method to identify tables containing lab testing results in scanned laboratory reports.
Methods
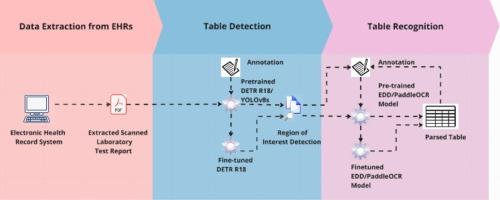
Extracting tabular data from scanned lab reports involves two stages: table detection (i.e., identifying the area of a table object) and table recognition (i.e., identifying and extracting tabular structures and contents). DETR R18 algorithm as well as YOLOv8s were involved for table detection, and we compared the performance of PaddleOCR and the encoder-dual-decoder (EDD) model for table recognition. 650 tables from 632 randomly selected laboratory test reports were annotated and used to train and evaluate those models. For table detection evaluation, we used metrics such as Average Precision (AP), Average Recall (AR), AP50, and AP75. For table recognition evaluation, we employed Tree-Edit Distance (TEDS).
Results
For table detection, fine-tuned DETR R18 demonstrated superior performance (AP50: 0.774; AP75: 0.644; AP: 0.601; AR: 0.766). In terms of table recognition, fine-tuned EDD outperformed other models with a TEDS score of 0.815. The proposed OCR pipeline (fine-tuned DETR R18 and fine-tuned EDD), demonstrated impressive results, achieving a TEDS score of 0.699 and a TEDS structure score of 0.764.
Conclusions
Our study presents a dedicated OCR pipeline for scanned clinical documents, utilizing state-of-the-art deep learning models for region-of-interest detection and table recognition. The high TEDS scores demonstrate the effectiveness of our approach, which has significant implications for clinical data analysis and decision-making.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Journal of Biomedical Informatics
医学-计算机:跨学科应用
CiteScore
8.90
自引率
6.70%
发文量
243
审稿时长
32 days
期刊介绍:
The Journal of Biomedical Informatics reflects a commitment to high-quality original research papers, reviews, and commentaries in the area of biomedical informatics methodology. Although we publish articles motivated by applications in the biomedical sciences (for example, clinical medicine, health care, population health, and translational bioinformatics), the journal emphasizes reports of new methodologies and techniques that have general applicability and that form the basis for the evolving science of biomedical informatics. Articles on medical devices; evaluations of implemented systems (including clinical trials of information technologies); or papers that provide insight into a biological process, a specific disease, or treatment options would generally be more suitable for publication in other venues. Papers on applications of signal processing and image analysis are often more suitable for biomedical engineering journals or other informatics journals, although we do publish papers that emphasize the information management and knowledge representation/modeling issues that arise in the storage and use of biological signals and images. System descriptions are welcome if they illustrate and substantiate the underlying methodology that is the principal focus of the report and an effort is made to address the generalizability and/or range of application of that methodology. Note also that, given the international nature of JBI, papers that deal with specific languages other than English, or with country-specific health systems or approaches, are acceptable for JBI only if they offer generalizable lessons that are relevant to the broad JBI readership, regardless of their country, language, culture, or health system.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


