利用通用领域资源增强生物医学命名实体识别。
IF 4
2区 医学
Q2 COMPUTER SCIENCE, INTERDISCIPLINARY APPLICATIONS
引用次数: 0
摘要
目的:训练基于神经网络的生物医学命名实体识别2(BioNER)模型通常需要大量昂贵的人工标注。虽然有几项研究利用多个 BioNER 数据集进行多任务学习以减少人力,但这种方法并不能持续提高性能,而且可能会在不同的生物医学语料库中引入标签模糊性。我们的目标是通过从与生物医学数据集概念重叠较少且易于获取的资源中进行迁移学习来应对这些挑战:我们提出了 GERBERA,一种利用通用领域 NER 数据集进行训练的简单而有效的方法。我们采用多任务学习法,利用目标 BioNER 数据集和通用域数据集训练一个预训练生物医学语言模型。随后,我们专门针对 BioNER 数据集对模型进行了微调:我们在八个实体类型的五个数据集上对 GERBERA 进行了系统评估,这些数据集共包含 81,410 个实例。尽管使用的生物医学资源较少,但与使用其他 BioNER 数据集训练的基线模型相比,我们的模型表现出了卓越的性能。具体来说,在八种实体类型中,我们的模型在六种类型中的表现始终优于基线模型,在八个实体中比最佳基线表现平均提高了 0.9%。我们的方法在数据有限的 BioNER 数据集上尤其有效,在 JNLPBA-RNA 数据集上的 F1 分数提高了 4.7%:本研究介绍了一种新的训练方法,该方法利用具有成本效益的通用领域 NER 数据集来增强 BioNER 模型。这种方法大大提高了 BioNER 模型的性能,使其成为生物医学数据集稀缺或昂贵情况下的宝贵资产。我们通过 https://github.com/qingyu-qc/bioner_gerbera 公开数据、代码和模型。本文章由计算机程序翻译,如有差异,请以英文原文为准。
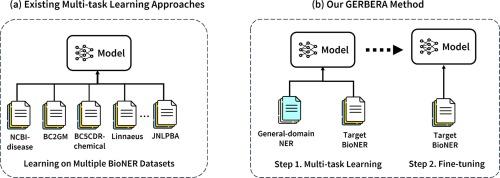
Augmenting biomedical named entity recognition with general-domain resources
Objective
Training a neural network-based biomedical named entity recognition (BioNER) model usually requires extensive and costly human annotations. While several studies have employed multi-task learning with multiple BioNER datasets to reduce human effort, this approach does not consistently yield performance improvements and may introduce label ambiguity in different biomedical corpora. We aim to tackle those challenges through transfer learning from easily accessible resources with fewer concept overlaps with biomedical datasets.
Methods
We proposed GERBERA, a simple-yet-effective method that utilized general-domain NER datasets for training. We performed multi-task learning to train a pre-trained biomedical language model with both the target BioNER dataset and the general-domain dataset. Subsequently, we fine-tuned the models specifically for the BioNER dataset.
Results
We systematically evaluated GERBERA on five datasets of eight entity types, collectively consisting of 81,410 instances. Despite using fewer biomedical resources, our models demonstrated superior performance compared to baseline models trained with additional BioNER datasets. Specifically, our models consistently outperformed the baseline models in six out of eight entity types, achieving an average improvement of 0.9% over the best baseline performance across eight entities. Our method was especially effective in amplifying performance on BioNER datasets characterized by limited data, with a 4.7% improvement in F1 scores on the JNLPBA-RNA dataset.
Conclusion
This study introduces a new training method that leverages cost-effective general-domain NER datasets to augment BioNER models. This approach significantly improves BioNER model performance, making it a valuable asset for scenarios with scarce or costly biomedical datasets. We make data, codes, and models publicly available via https://github.com/qingyu-qc/bioner_gerbera.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Journal of Biomedical Informatics
医学-计算机:跨学科应用
CiteScore
8.90
自引率
6.70%
发文量
243
审稿时长
32 days
期刊介绍:
The Journal of Biomedical Informatics reflects a commitment to high-quality original research papers, reviews, and commentaries in the area of biomedical informatics methodology. Although we publish articles motivated by applications in the biomedical sciences (for example, clinical medicine, health care, population health, and translational bioinformatics), the journal emphasizes reports of new methodologies and techniques that have general applicability and that form the basis for the evolving science of biomedical informatics. Articles on medical devices; evaluations of implemented systems (including clinical trials of information technologies); or papers that provide insight into a biological process, a specific disease, or treatment options would generally be more suitable for publication in other venues. Papers on applications of signal processing and image analysis are often more suitable for biomedical engineering journals or other informatics journals, although we do publish papers that emphasize the information management and knowledge representation/modeling issues that arise in the storage and use of biological signals and images. System descriptions are welcome if they illustrate and substantiate the underlying methodology that is the principal focus of the report and an effort is made to address the generalizability and/or range of application of that methodology. Note also that, given the international nature of JBI, papers that deal with specific languages other than English, or with country-specific health systems or approaches, are acceptable for JBI only if they offer generalizable lessons that are relevant to the broad JBI readership, regardless of their country, language, culture, or health system.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


