人类性腺性别决定布尔模型的动态稳健性。
IF 3.1
4区 生物学
Q2 BIOLOGY
引用次数: 0
摘要
性腺性别决定(GSD)是胚胎发育早期的一个复杂过程,但人们对其了解甚少。这一过程分别通过激活与 Sertoli 或 Granulosa 细胞相关的遗传因子,决定双潜能性腺原基(BGP)是分化成睾丸还是卵巢。由于实验的局限性和潜在基因相互作用的复杂性,对这一发育过程的研究仍具有挑战性。布尔网络(BN)是模拟遗传行为的二元网络,由于其在处理大量基因相互作用时的简便性,常用于基因调控网络(GRN)的建模。已报道的二元网络通常使用同步(并行)更新方案,即所有节点(代表基因)同时更新其值。然而,使用这种更新方案受到了批评,因为它无法表示在时间尺度上受到高度调控的生物系统。异步更新方案和块序列更新方案是解决这一问题的替代方案。在第一种情况下,更新方案遵循随机行为,而在第二种情况下,网络节点集被划分为若干区块,这样一个区块内的节点会同时更新,而且这些区块是按照特定顺序依次考虑的。为了评估不同更新方法对与 GSD 相关的 GRN 的影响,我们首先在不丢失原始网络主要动态(与睾丸和卵巢的形成相关)的情况下减少了节点。然后,我们测试了基因失活所产生的扰动对网络吸引子的影响,特别是 SRY 和 WNT4 基因,因为前者只存在于 Y 染色体中,而后者在早期胚胎发育中非常重要。我们发现这两个基因都很重要,但 WNT4 单独显示的表型吸引子比例要高于 SRY 单独显示的吸引子比例。最后,我们发现使用异步和块序列更新方案时,吸引盆地(即达到吸引子的配置集)的百分比与原始网络的百分比相似,这支持了模型的鲁棒性。本文章由计算机程序翻译,如有差异,请以英文原文为准。
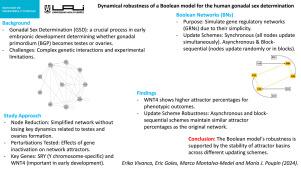
Dynamical robustness of a Boolean model for the human gonadal sex determination
Gonadal sex determination (GSD) is a complex but poorly understood process in the early stages of embryonic development. This process determines whether the bipotential gonadal primordium (BGP) will differentiate into testes or ovaries through the activation of genetic factors related to Sertoli or Granulosa cells, respectively. The study of this developmental process remains challenging due to experimental limitations and the complexity of the underlying genetic interactions. Boolean Networks (BNs) are binary networks that simulate genetic behavior and are commonly used for modeling gene regulatory networks (GRNs) due to their simplicity when dealing with a high number of gene interactions. Reported BNs usually use a synchronous (parallel) update scheme, which means that all the nodes (representing genes) update their values simultaneously. However, the use of this update scheme has been criticized because it cannot represent biological systems that are highly regulated at a temporal scale. Asynchronous and block-sequential updating schemes appear as an alternative to tackle this issue. In the first case, the updating scheme follows a random behavior while, in the second case, the set of network nodes is partitioned into blocks such that the nodes within a block are updated simultaneously, and the blocks are considered in a specific order sequence. To assess the impact of different updating approaches in a GRN associated to GSD we first made a node reduction without losing the main dynamics of the original network which are related to the formation of testes and ovaries. Then, we tested the effect of perturbations given by the inactivation of genes on the network attractors, specifically the SRY and WNT4 genes, since the former is only present in the Y chromosome and the latter is of importance in early embryo development. We found that both genes were crucial, but WNT4 alone showed a higher percentage of attractors towards a phenotype than the SRY alone. Finally, we found that using asynchronous and block-sequential updating schemes, the attraction basins – i.e., the set of configurations that reach an attractor – remain with similar percentages to those of the original network, which supports the robustness of the model.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Computational Biology and Chemistry
生物-计算机:跨学科应用
CiteScore
6.10
自引率
3.20%
发文量
142
审稿时长
24 days
期刊介绍:
Computational Biology and Chemistry publishes original research papers and review articles in all areas of computational life sciences. High quality research contributions with a major computational component in the areas of nucleic acid and protein sequence research, molecular evolution, molecular genetics (functional genomics and proteomics), theory and practice of either biology-specific or chemical-biology-specific modeling, and structural biology of nucleic acids and proteins are particularly welcome. Exceptionally high quality research work in bioinformatics, systems biology, ecology, computational pharmacology, metabolism, biomedical engineering, epidemiology, and statistical genetics will also be considered.
Given their inherent uncertainty, protein modeling and molecular docking studies should be thoroughly validated. In the absence of experimental results for validation, the use of molecular dynamics simulations along with detailed free energy calculations, for example, should be used as complementary techniques to support the major conclusions. Submissions of premature modeling exercises without additional biological insights will not be considered.
Review articles will generally be commissioned by the editors and should not be submitted to the journal without explicit invitation. However prospective authors are welcome to send a brief (one to three pages) synopsis, which will be evaluated by the editors.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


