开发基于机器学习的靶标特异性评分功能,为人类二氢烟酸脱氢酶抑制剂进行基于结构的结合亲和力预测
IF 3.4
3区 化学
Q2 CHEMISTRY, MULTIDISCIPLINARY
引用次数: 0
摘要
人二氢烟酸脱氢酶(hDHODH)是一种依赖于黄素单核苷酸的酶,可以限制嘧啶的从头合成,使其成为自身免疫性疾病和癌症等疾病的治疗靶点。本研究利用 AutoDock Vina 生成的复合物对接结构,整合了相互作用特征和配体特征,并采用支持向量回归开发了 hDHODH 的靶标特异性评分函数(TSSF-hDHODH)。在交叉验证和外部验证中,TSSF-hDHODH 的皮尔逊相关系数分别为 0.86 和 0.74,均远远优于经典评分函数 AutoDock Vina 和基于随机森林(RF)的通用评分函数 RF-Score。TSSF-hDHODH 还被进一步用于虚拟筛选 FDA 批准的药典药物库中的潜在抑制剂。结合分子动力学模拟的结果,克唑替尼被确定为后续结构优化的候选药物。这项研究有助于发现 hDHODH 抑制剂,并为其他靶点开发评分功能。本文章由计算机程序翻译,如有差异,请以英文原文为准。
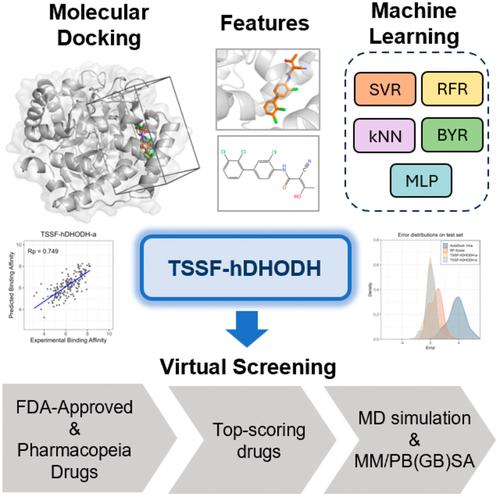
Development of a machine learning-based target-specific scoring function for structure-based binding affinity prediction for human dihydroorotate dehydrogenase inhibitors
Human dihydroorotate dehydrogenase (hDHODH) is a flavin mononucleotide-dependent enzyme that can limit de novo pyrimidine synthesis, making it a therapeutic target for diseases such as autoimmune disorders and cancer. In this study, using the docking structures of complexes generated by AutoDock Vina, we integrate interaction features and ligand features, and employ support vector regression to develop a target-specific scoring function for hDHODH (TSSF-hDHODH). The Pearson correlation coefficient values of TSSF-hDHODH in the cross-validation and external validation are 0.86 and 0.74, respectively, both of which are far superior to those of classic scoring function AutoDock Vina and random forest (RF) based generic scoring function RF-Score. TSSF-hDHODH is further used for the virtual screening of potential inhibitors in the FDA-Approved & Pharmacopeia Drug Library. In conjunction with the results from molecular dynamics simulations, crizotinib is identified as a candidate for subsequent structural optimization. This study can be useful for the discovery of hDHODH inhibitors and the development of scoring functions for additional targets.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
CiteScore
6.60
自引率
3.30%
发文量
247
审稿时长
1.7 months
期刊介绍:
This distinguished journal publishes articles concerned with all aspects of computational chemistry: analytical, biological, inorganic, organic, physical, and materials. The Journal of Computational Chemistry presents original research, contemporary developments in theory and methodology, and state-of-the-art applications. Computational areas that are featured in the journal include ab initio and semiempirical quantum mechanics, density functional theory, molecular mechanics, molecular dynamics, statistical mechanics, cheminformatics, biomolecular structure prediction, molecular design, and bioinformatics.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


