组织病理学图像的再识别
IF 10.7
1区 医学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
摘要
在大量研究中,深度学习算法证明了其在组织病理学图像分析方面的潜力,例如,用于揭示肿瘤的亚型或转移瘤的原发来源。这些模型需要大量数据集进行训练,这些数据集必须进行匿名处理,以防止可能出现的患者身份泄露。本研究证明,即使是相对简单的深度学习算法,也能在大型组织病理学数据集中重新识别患者,而且准确率很高。此外,我们还针对给定任务比较了一整套最先进的全切片图像分类器和特征提取器。我们在两个 TCIA 数据集上评估了我们的算法,包括肺鳞状细胞癌(LSCC)和肺腺癌(LUAD)。我们还展示了算法在脑膜瘤组织内部数据集上的性能。在 LSCC 和 LUAD 数据集上,我们预测玻片来源患者的 F1 分数分别高达 80.1% 和 77.19%,而在脑膜瘤数据集上则高达 77.09%。根据我们的研究结果,我们制定了一个风险评估方案,以估算发表前患者隐私所面临的风险。本文章由计算机程序翻译,如有差异,请以英文原文为准。
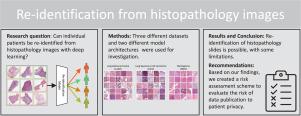
Re-identification from histopathology images
In numerous studies, deep learning algorithms have proven their potential for the analysis of histopathology images, for example, for revealing the subtypes of tumors or the primary origin of metastases. These models require large datasets for training, which must be anonymized to prevent possible patient identity leaks. This study demonstrates that even relatively simple deep learning algorithms can re-identify patients in large histopathology datasets with substantial accuracy. In addition, we compared a comprehensive set of state-of-the-art whole slide image classifiers and feature extractors for the given task. We evaluated our algorithms on two TCIA datasets including lung squamous cell carcinoma (LSCC) and lung adenocarcinoma (LUAD). We also demonstrate the algorithm’s performance on an in-house dataset of meningioma tissue. We predicted the source patient of a slide with scores of up to 80.1% and 77.19% on the LSCC and LUAD datasets, respectively, and with 77.09% on our meningioma dataset. Based on our findings, we formulated a risk assessment scheme to estimate the risk to the patient’s privacy prior to publication.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Medical image analysis
工程技术-工程:生物医学
CiteScore
22.10
自引率
6.40%
发文量
309
审稿时长
6.6 months
期刊介绍:
Medical Image Analysis serves as a platform for sharing new research findings in the realm of medical and biological image analysis, with a focus on applications of computer vision, virtual reality, and robotics to biomedical imaging challenges. The journal prioritizes the publication of high-quality, original papers contributing to the fundamental science of processing, analyzing, and utilizing medical and biological images. It welcomes approaches utilizing biomedical image datasets across all spatial scales, from molecular/cellular imaging to tissue/organ imaging.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


