用于结构化医疗信息检索的隐私保护大型语言模型。
IF 12.4
1区 医学
Q1 HEALTH CARE SCIENCES & SERVICES
引用次数: 0
摘要
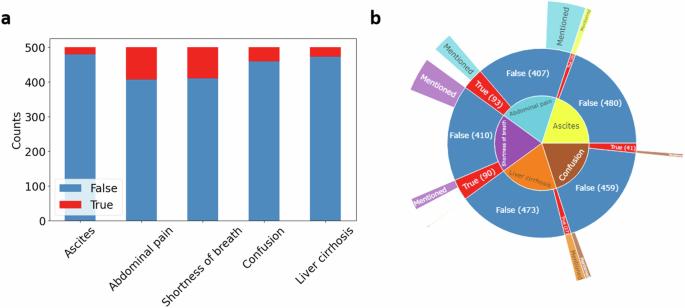
大多数临床信息以自由文本形式编码,无法进行定量分析。本研究提出了一个开源管道,使用本地大语言模型(LLM)"Llama 2 "从临床文本中提取定量信息,并评估了其在识别肝硬化失代偿期特征方面的性能。LLM 从 MIMIC IV 数据集中的 500 份患者病历中,以零点和单点的方式识别出了五个关键临床特征。我们比较了三种规模的 LLM 和各种提示工程方法,并将预测结果与三位盲人医学专家提供的地面实况进行了比较。我们的管道具有很高的准确性,检测肝硬化的灵敏度为 100%,特异性为 96%。使用 700 亿参数模型检测腹水(95%,95%)、混淆(76%,94%)、腹痛(84%,97%)和呼吸急促(87%,97%)时,灵敏度和特异性也很高,其表现优于较小的版本。我们的研究成功证明了本地部署的 LLM 能够以较低的硬件要求从自由文本中提取临床信息。本文章由计算机程序翻译,如有差异,请以英文原文为准。
Privacy-preserving large language models for structured medical information retrieval
Most clinical information is encoded as free text, not accessible for quantitative analysis. This study presents an open-source pipeline using the local large language model (LLM) “Llama 2” to extract quantitative information from clinical text and evaluates its performance in identifying features of decompensated liver cirrhosis. The LLM identified five key clinical features in a zero- and one-shot manner from 500 patient medical histories in the MIMIC IV dataset. We compared LLMs of three sizes and various prompt engineering approaches, with predictions compared against ground truth from three blinded medical experts. Our pipeline achieved high accuracy, detecting liver cirrhosis with 100% sensitivity and 96% specificity. High sensitivities and specificities were also yielded for detecting ascites (95%, 95%), confusion (76%, 94%), abdominal pain (84%, 97%), and shortness of breath (87%, 97%) using the 70 billion parameter model, which outperformed smaller versions. Our study successfully demonstrates the capability of locally deployed LLMs to extract clinical information from free text with low hardware requirements.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

NPJ Digital Medicine
Multiple-
CiteScore
25.10
自引率
3.30%
发文量
170
审稿时长
15 weeks
期刊介绍:
npj Digital Medicine is an online open-access journal that focuses on publishing peer-reviewed research in the field of digital medicine. The journal covers various aspects of digital medicine, including the application and implementation of digital and mobile technologies in clinical settings, virtual healthcare, and the use of artificial intelligence and informatics.
The primary goal of the journal is to support innovation and the advancement of healthcare through the integration of new digital and mobile technologies. When determining if a manuscript is suitable for publication, the journal considers four important criteria: novelty, clinical relevance, scientific rigor, and digital innovation.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


