用于化学蛋白质组学标记和靶向分析的 BODIPY 嵌入式同位素特征的研究与开发
IF 3.1
2区 化学
Q2 BIOCHEMICAL RESEARCH METHODS
Journal of the American Society for Mass Spectrometry
Pub Date : 2024-09-16
DOI:10.1021/jasms.4c00246
引用次数: 0
摘要
基于质谱的化学蛋白质组学的一个共同目标是直接测量目标蛋白质与小分子配体之间的连接位点。然而,由于标记蛋白质的丰度较低,而且使用标准蛋白质组学软件难以确定修饰位点,因此这些实验本身就具有挑战性。产生特征性片段离子或同位素编码目标肽的报告标签可用于预先发现标记肽,即使在没有鉴定的情况下也是如此。我们研究了 BODIPY FL 叠氮化物作为化学蛋白质组学试剂的潜力,因为它含有硼,而且 10B:11B 的天然丰度比为 1:4。硼的同位素编码 BODIPY 标记的肽,在单异位(M)峰和 M+1 峰之间有可预测的模式。使用 R 脚本在 MS1 图谱中识别 BODIPY 标记的肽段,该脚本可过滤特征性的 10B:11B 强度比和质量缺陷。应用硼检测脚本使 BODIPY 结合的 BSA 样品的标记肽覆盖率是非目标数据依赖性采集测序的三倍。此外,我们还利用 BODIPY 固有的高频中性丢失特征来协助 BODIPY 修饰肽的鉴定。最后,我们使用添加到复杂大肠杆菌消化液中的 BODIPY 结合物 BSA 样品演示了这种方法的应用。总之,我们的研究结果表明,将市售的 BODIPY FL 叠氮化物与炔烃标记的肽相连接,可提供一种独特的同位素特征,用于精确定位修饰位点,并可进行在线或离线紫外或荧光检测。本文章由计算机程序翻译,如有差异,请以英文原文为准。
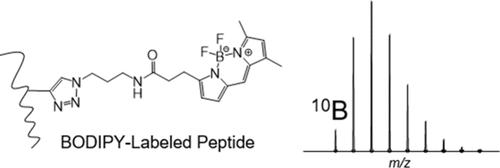
Investigation and Development of the BODIPY-Embedded Isotopic Signature for Chemoproteomics Labeling and Targeted Profiling
A common goal in mass spectrometry-based chemoproteomics is to directly measure the site of conjugation between the target protein and the small molecule ligand. However, these experiments are inherently challenging due to the low abundance of labeled proteins and the difficulty in identifying modification sites using standard proteomics software. Reporter tags that either generate signature fragment ions or isotopically encode target peptides can be used for the preemptive discovery of labeled peptides even in the absence of identification. We investigated the potential of BODIPY FL azide as a click chemistry enabled chemoproteomics reagent due to the presence of boron and the unique 1:4 natural abundance ratio of 10B:11B. The isotopes of boron encode BODIPY-labeled peptides with a predictable pattern between the monoisotopic (M) and M+1 peaks. BODIPY-labeled peptides were identified in MS1 spectra using an R script that filters for the signature 10B:11B intensity ratio and mass defect. Application of the boron detection script resulted in three times the labeled peptide coverage achieved for a BODIPY-conjugated BSA sample compared with untargeted data-dependent acquisition sequencing. Furthermore, we used the inherent HF neutral loss signature from BODIPY to assist with BODIPY-modified peptide identification. Finally, we demonstrate the application of this approach using the BODIPY-conjugated BSA sample spiked into a complex E. coli. digest. In summary, our results show that the commercially available BODIPY FL azide clicked to alkyne-labeled peptides provides a unique isotopic signature for pinpointing the site(s) of modification with the added potential for on- or off-line UV or fluorescence detection.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
CiteScore
5.50
自引率
9.40%
发文量
257
审稿时长
1 months
期刊介绍:
The Journal of the American Society for Mass Spectrometry presents research papers covering all aspects of mass spectrometry, incorporating coverage of fields of scientific inquiry in which mass spectrometry can play a role.
Comprehensive in scope, the journal publishes papers on both fundamentals and applications of mass spectrometry. Fundamental subjects include instrumentation principles, design, and demonstration, structures and chemical properties of gas-phase ions, studies of thermodynamic properties, ion spectroscopy, chemical kinetics, mechanisms of ionization, theories of ion fragmentation, cluster ions, and potential energy surfaces. In addition to full papers, the journal offers Communications, Application Notes, and Accounts and Perspectives

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


