利用 DiffPepBuilder 进行靶标特异性新肽粘合剂设计
IF 5.6
2区 化学
Q1 CHEMISTRY, MEDICINAL
引用次数: 0
摘要
尽管在靶标特异性从头蛋白质粘合剂设计方面取得了令人振奋的进展,但由于多肽结构的灵活性和蛋白质-多肽复合结构数据的稀缺性,多肽粘合剂设计仍然具有挑战性。在这项研究中,我们从丰富的蛋白质-蛋白质界面数据中整理出了一个大型合成数据集(称为 PepPC-F),并开发出了 DiffPepBuilder,这是一种全新的靶标特异性多肽粘合剂生成方法,它利用在 PepPC-F 上训练的 SE(3)-equivariant diffusion 模型来编码设计多肽序列和结构。DiffPepBuilder 还引入了二硫键来稳定生成的多肽结构。我们在 30 个经实验验证的强肽结合体上测试了 DiffPepBuilder,这些结合体具有可用的蛋白质-肽复合物结构。DiffPepBuilder 能够有效地重现多肽配体的原生结构和序列,并生成具有更高的结合自由能的新型多肽结合体。随后,我们对三个目标进行了从头生成案例研究。在再生测试和案例研究中,DiffPepBuilder 在序列和结构回忆、界面质量和结构多样性方面都优于 AfDesign 和 RFdiffusion 以及 ProteinMPNN。分子动力学模拟证实,二硫键的引入增强了生成多肽的结构刚性和结合性能。作为一种通用的多肽结合剂从头设计工具,DiffPepBuilder 可用于根据三维和结合位点信息为给定的蛋白质目标设计多肽结合剂。本文章由计算机程序翻译,如有差异,请以英文原文为准。
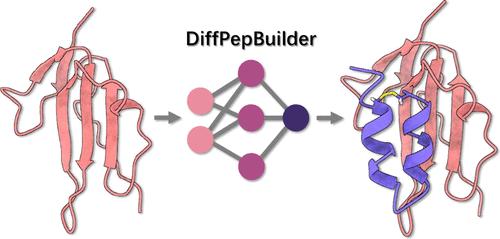
Target-Specific De Novo Peptide Binder Design with DiffPepBuilder
Despite the exciting progress in target-specific de novo protein binder design, peptide binder design remains challenging due to the flexibility of peptide structures and the scarcity of protein-peptide complex structure data. In this study, we curated a large synthetic data set, referred to as PepPC-F, from the abundant protein–protein interface data and developed DiffPepBuilder, a de novo target-specific peptide binder generation method that utilizes an SE(3)-equivariant diffusion model trained on PepPC-F to codesign peptide sequences and structures. DiffPepBuilder also introduces disulfide bonds to stabilize the generated peptide structures. We tested DiffPepBuilder on 30 experimentally verified strong peptide binders with available protein–peptide complex structures. DiffPepBuilder was able to effectively recall the native structures and sequences of the peptide ligands and to generate novel peptide binders with improved binding free energy. We subsequently conducted de novo generation case studies on three targets. In both the regeneration test and case studies, DiffPepBuilder outperformed AfDesign and RFdiffusion coupled with ProteinMPNN, in terms of sequence and structure recall, interface quality, and structural diversity. Molecular dynamics simulations confirmed that the introduction of disulfide bonds enhanced the structural rigidity and binding performance of the generated peptides. As a general peptide binder de novo design tool, DiffPepBuilder can be used to design peptide binders for given protein targets with three-dimensional and binding site information.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊
CiteScore
9.80
自引率
10.70%
发文量
529
审稿时长
1.4 months
期刊介绍:
The Journal of Chemical Information and Modeling publishes papers reporting new methodology and/or important applications in the fields of chemical informatics and molecular modeling. Specific topics include the representation and computer-based searching of chemical databases, molecular modeling, computer-aided molecular design of new materials, catalysts, or ligands, development of new computational methods or efficient algorithms for chemical software, and biopharmaceutical chemistry including analyses of biological activity and other issues related to drug discovery.
Astute chemists, computer scientists, and information specialists look to this monthly’s insightful research studies, programming innovations, and software reviews to keep current with advances in this integral, multidisciplinary field.
As a subscriber you’ll stay abreast of database search systems, use of graph theory in chemical problems, substructure search systems, pattern recognition and clustering, analysis of chemical and physical data, molecular modeling, graphics and natural language interfaces, bibliometric and citation analysis, and synthesis design and reactions databases.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


