T 细胞急性淋巴细胞白血病对 SWI/SNF 染色质重塑因子的致癌依赖性。
IF 12.8
1区 医学
Q1 HEMATOLOGY
引用次数: 0
摘要
T 细胞急性淋巴细胞白血病(T-ALL)是一种由未成熟胸腺细胞引起的血液恶性肿瘤。与 NOTCH1 和 MYC 等众所周知的致癌转录因子不同,人们对染色质重塑因子参与 T-ALL 发病机制的情况知之甚少。在此,我们提供了令人信服的证据,证明 SWI/SNF 染色质重塑复合物如何参与人类 T-ALL 发病机制。转录组和 ATAC-Seq 数据集的整合分析表明,与正常 T 细胞相比,SWI/SNF 复合物亚基之一 SMARCA4 在 T-ALL 患者样本和细胞系中的高表达。SMARCA 蛋白功能缺失导致多种 T-ALL 细胞系凋亡诱导和生长抑制。ATAC-Seq 分析显示,在 SMARCA 蛋白功能缺失后,整个基因组的染色质可及性大幅降低。RUNX1 与 SMARCA4 蛋白相互作用,共同占据相同的基因组区域。重要的是,当SMARCA蛋白功能受损时,NOTCH1-MYC通路主要受到影响,这表明SWI/SNF是一个新的治疗靶点。本文章由计算机程序翻译,如有差异,请以英文原文为准。
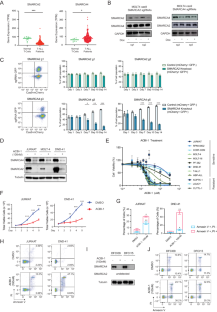
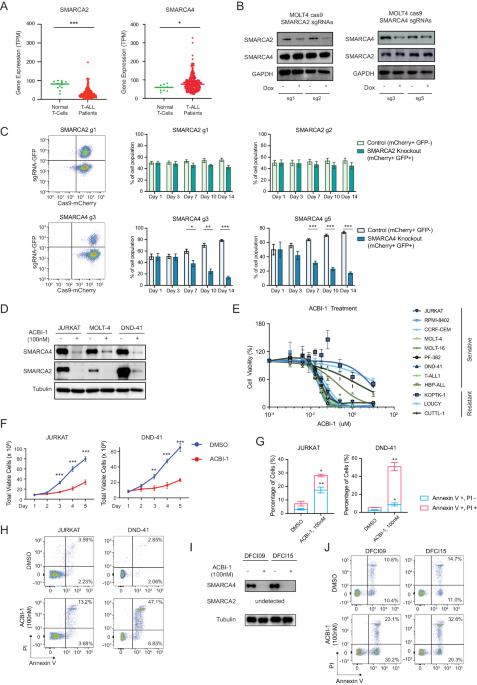
Oncogenic dependency on SWI/SNF chromatin remodeling factors in T-cell acute lymphoblastic leukemia
T-cell acute lymphoblastic leukemia (T-ALL) is a hematological malignancy arising from immature thymocytes. Unlike well-known oncogenic transcription factors, such as NOTCH1 and MYC, the involvement of chromatin remodeling factors in T-ALL pathogenesis is poorly understood. Here, we provide compelling evidence on how SWI/SNF chromatin remodeling complex contributes to human T-ALL pathogenesis. Integrative analysis of transcriptomic and ATAC-Seq datasets revealed high expression of SMARCA4, one of the subunits of the SWI/SNF complex, in T-ALL patient samples and cell lines compared to normal T cells. Loss of SMARCA protein function resulted in apoptosis induction and growth inhibition in multiple T-ALL cell lines. ATAC-Seq analysis revealed a massive reduction in chromatin accessibility across the genome after the loss of SMARCA protein function. RUNX1 interacts with SMARCA4 protein and co-occupies the same genomic regions. Importantly, the NOTCH1-MYC pathway was primarily affected when SMARCA protein function was impaired, implicating SWI/SNF as a novel therapeutic target.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Leukemia
医学-血液学
CiteScore
18.10
自引率
3.50%
发文量
270
审稿时长
3-6 weeks
期刊介绍:
Title: Leukemia
Journal Overview:
Publishes high-quality, peer-reviewed research
Covers all aspects of research and treatment of leukemia and allied diseases
Includes studies of normal hemopoiesis due to comparative relevance
Topics of Interest:
Oncogenes
Growth factors
Stem cells
Leukemia genomics
Cell cycle
Signal transduction
Molecular targets for therapy
And more
Content Types:
Original research articles
Reviews
Letters
Correspondence
Comments elaborating on significant advances and covering topical issues
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


