用于蛋白质配体相互作用预测的多尺度拓扑结构-序列转换器
IF 18.8
1区 计算机科学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
摘要
尽管预训练自然语言处理(NLP)模型在各个领域都取得了成功,但它们在计算生物学中的应用却因依赖生物序列而受到阻碍,因为生物序列忽略了重要的三维(3D)结构信息,与 NLP 模型的序列架构不兼容。在这里,我们介绍一种拓扑变换器(TopoFormer),它是通过整合 NLP 模型和多尺度拓扑技术--持久拓扑超图拉普拉斯(PTHL)--而建立起来的,它能系统地将各种空间尺度上错综复杂的三维蛋白质配体复合物转换成 NLP 允许的拓扑不变式和同位形状序列。PTHL 将复杂的三维蛋白质配体复合物系统地转换成与 NLP 兼容的拓扑不变式和形状序列,捕捉跨空间尺度的基本相互作用。TopoFormer 在多个基准数据集的排序、对接和筛选任务中,具有极高的评分准确性和出色的性能。这种方法可用于将一般的高维结构化数据转换成与 NLP 兼容的序列,为更广泛的基于 NLP 的研究铺平道路。本文章由计算机程序翻译,如有差异,请以英文原文为准。
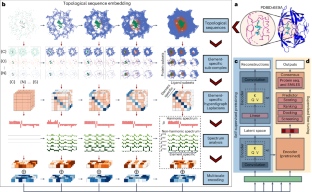
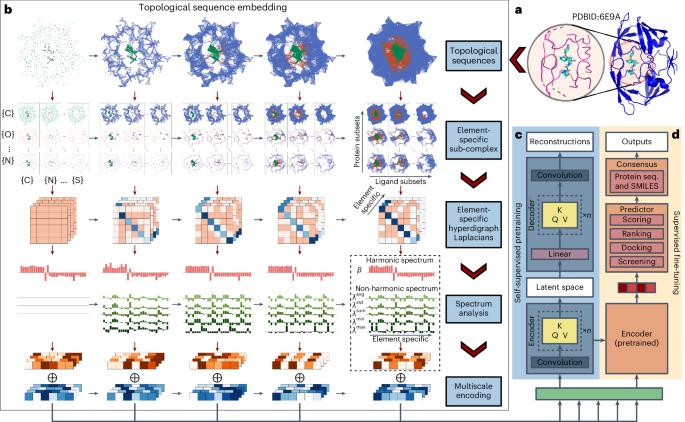
Multiscale topology-enabled structure-to-sequence transformer for protein–ligand interaction predictions
Despite the success of pretrained natural language processing (NLP) models in various fields, their application in computational biology has been hindered by their reliance on biological sequences, which ignores vital three-dimensional (3D) structural information incompatible with the sequential architecture of NLP models. Here we present a topological transformer (TopoFormer), which is built by integrating NLP models and a multiscale topology technique, the persistent topological hyperdigraph Laplacian (PTHL), which systematically converts intricate 3D protein–ligand complexes at various spatial scales into an NLP-admissible sequence of topological invariants and homotopic shapes. PTHL systematically transforms intricate 3D protein–ligand complexes into NLP-compatible sequences of topological invariants and shapes, capturing essential interactions across spatial scales. TopoFormer gives rise to exemplary scoring accuracy and excellent performance in ranking, docking and screening tasks in several benchmark datasets. This approach can be utilized to convert general high-dimensional structured data into NLP-compatible sequences, paving the way for broader NLP based research. Transformers show much promise for applications in computational biology, but they rely on sequences, and a challenge is to incorporate 3D structural information. TopoFormer, proposed by Dong Chen et al., combines transformers with a mathematical multiscale topology technique to model 3D protein–ligand complexes, substantially enhancing performance in a range of prediction tasks of interest to drug discovery.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Machine Intelligence
Multiple-
CiteScore
36.90
自引率
2.10%
发文量
127
期刊介绍:
Nature Machine Intelligence is a distinguished publication that presents original research and reviews on various topics in machine learning, robotics, and AI. Our focus extends beyond these fields, exploring their profound impact on other scientific disciplines, as well as societal and industrial aspects. We recognize limitless possibilities wherein machine intelligence can augment human capabilities and knowledge in domains like scientific exploration, healthcare, medical diagnostics, and the creation of safe and sustainable cities, transportation, and agriculture. Simultaneously, we acknowledge the emergence of ethical, social, and legal concerns due to the rapid pace of advancements.
To foster interdisciplinary discussions on these far-reaching implications, Nature Machine Intelligence serves as a platform for dialogue facilitated through Comments, News Features, News & Views articles, and Correspondence. Our goal is to encourage a comprehensive examination of these subjects.
Similar to all Nature-branded journals, Nature Machine Intelligence operates under the guidance of a team of skilled editors. We adhere to a fair and rigorous peer-review process, ensuring high standards of copy-editing and production, swift publication, and editorial independence.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


