Protein–protein contact prediction by geometric triangle-aware protein language models
IF 23.9
1区 计算机科学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
Abstract
Information regarding the residue–residue distance between interacting proteins is important for modelling the structures of protein complexes, as well as being valuable for understanding the molecular mechanism of protein–protein interactions. With the advent of deep learning, many methods have been developed to accurately predict the intra-protein residue–residue contacts of monomers. However, it is still challenging to accurately predict inter-protein residue–residue contacts for protein complexes, especially hetero-protein complexes. Here we develop a protein language model-based deep learning method to predict the inter-protein residue–residue contacts of protein complexes—named DeepInter—by introducing a triangle-aware mechanism of triangle update and triangle self-attention into the deep neural network. We extensively validate DeepInter on diverse test sets of 300 homodimeric, 28 CASP-CAPRI homodimeric and 99 heterodimeric complexes and compare it with state-of-the-art methods including CDPred, DeepHomo2.0, GLINTER and DeepHomo. The results demonstrate the accuracy and robustness of DeepInter. Contact prediction between two proteins is still computationally challenging, but is vital for understanding multi-protein complexes. Lin et al. use a geometric deep learning approach to provide accurate predictions of inter-protein residue–residue contacts.
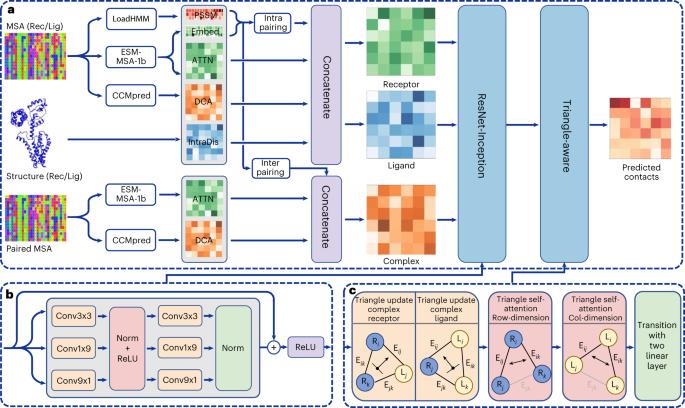
通过几何三角形感知蛋白质语言模型预测蛋白质与蛋白质之间的接触
有关相互作用蛋白质之间残基-残基距离的信息对于蛋白质复合物结构建模非常重要,对于理解蛋白质-蛋白质相互作用的分子机制也很有价值。随着深度学习技术的出现,人们已经开发出许多方法来准确预测单体蛋白质内部残基-残基接触。然而,准确预测蛋白质复合物(尤其是异源蛋白质复合物)的蛋白质残基-残基间接触仍具有挑战性。在此,我们开发了一种基于蛋白质语言模型的深度学习方法来预测蛋白质复合物的蛋白质残基-残基间的接触--命名为DeepInter--通过在深度神经网络中引入三角形更新和三角形自我关注的三角形感知机制。我们在300个同源二聚体、28个CASP-CAPRI同源二聚体和99个异源二聚体复合物的不同测试集上广泛验证了DeepInter,并将其与CDPred、DeepHomo2.0、GLINTER和DeepHomo等最先进的方法进行了比较。结果证明了 DeepInter 的准确性和稳健性。两个蛋白质之间的接触预测在计算上仍然具有挑战性,但对理解多蛋白复合物至关重要。Lin 等人使用几何深度学习方法对蛋白质残基-残基间的接触进行了准确预测。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Machine Intelligence
Multiple-
CiteScore
36.90
自引率
2.10%
发文量
127
期刊介绍:
Nature Machine Intelligence is a distinguished publication that presents original research and reviews on various topics in machine learning, robotics, and AI. Our focus extends beyond these fields, exploring their profound impact on other scientific disciplines, as well as societal and industrial aspects. We recognize limitless possibilities wherein machine intelligence can augment human capabilities and knowledge in domains like scientific exploration, healthcare, medical diagnostics, and the creation of safe and sustainable cities, transportation, and agriculture. Simultaneously, we acknowledge the emergence of ethical, social, and legal concerns due to the rapid pace of advancements.
To foster interdisciplinary discussions on these far-reaching implications, Nature Machine Intelligence serves as a platform for dialogue facilitated through Comments, News Features, News & Views articles, and Correspondence. Our goal is to encourage a comprehensive examination of these subjects.
Similar to all Nature-branded journals, Nature Machine Intelligence operates under the guidance of a team of skilled editors. We adhere to a fair and rigorous peer-review process, ensuring high standards of copy-editing and production, swift publication, and editorial independence.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


