Degradation of circular RNA by the ribonuclease DIS3
IF 14.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
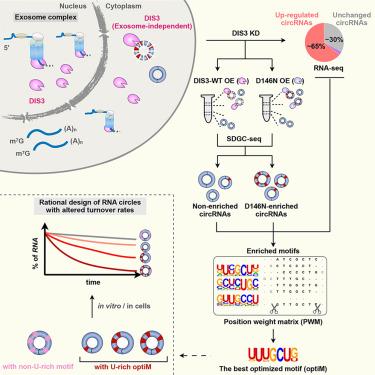
Features of circular RNAs (circRNAs) produced by back-splicing of eukaryotic exon(s) make them resistant to degradation by linear RNA decay machineries. Thus, a general circRNA degradation pathway under normal conditions has remained largely elusive. Here, we report that the endonucleolytic enzyme DIS3 is responsible for the degradation of circRNAs. Depletion of DIS3 leads to the upregulation of more than 60% of circRNAs with little effect on their linear cognates. Such DIS3-mediated circRNA degradation is conserved, occurs in the cytoplasm, and relies on DIS3’s endonucleolytic activity but is independent of the RNA exosome complex. Sequence enrichment analyses suggest that DIS3 prefers to degrade circRNAs containing U-rich motifs. Correspondingly, synthesized RNA circles with or without U-rich motifs exhibit decreased or increased stabilities, respectively. Together, these findings suggest a general regulation of circRNA turnover by DIS3.
求助全文
约1分钟内获得全文
求助全文
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


