Analogue-Sensitive Inhibition of Histone Demethylases Uncovers Member-Specific Function in Ribosomal Protein Synthesis
IF 14.4
1区 化学
Q1 CHEMISTRY, MULTIDISCIPLINARY
引用次数: 0
Abstract
Lysine demethylases (KDMs) catalyze the oxidative removal of the methyl group from histones using earth-abundant iron and the metabolite 2-oxoglutarate (2OG). KDMs have emerged as master regulators of eukaryotic gene expression and are novel drug targets; small-molecule inhibitors of KDMs are in the clinical pipeline for the treatment of human cancer. Yet, mechanistic insights into the functional heterogeneity of human KDMs are limited, necessitating the development of chemical probes for precision targeting. Herein, we identify analogue-sensitive (as) mutants of the KDM4 subfamily to elucidate member-specific biological functions in a temporally defined manner. By replacing the highly conserved phenylalanine residue in the active site of KDM4 members with alanine, we develop mutants with intact catalytic activity and substrate specificity indistinguishable from those of the wild type congener. Unlike the wild type demethylases, mutants were sensitized toward cofactor-competitive N-oxalyl glycine (NOG) analogues carrying complementary steric appendage. Particularly notable is N-oxalyl leucine (NOL) which inhibited the KDM4 mutants reversibly with submicromolar efficacy. Cell-permeable NOL prodrugs inhibited as enzymes in cultured human cells to modulate lysine methylation on nucleosomal histones. Through conditional perturbation of the orthogonal enzymes, we uncover a KDM4A-specific role in ribosomal protein synthesis and map a remarkably dynamic signaling cascade involving locus-specific histone demethylation leading to fast rRNA expression, enhanced ribosome assembly, and protein synthesis. The results provide a mechanistic clue into KDM4A’s role in cancers that rely on heightened ribosomal activity to support uncontrolled cellular proliferation.
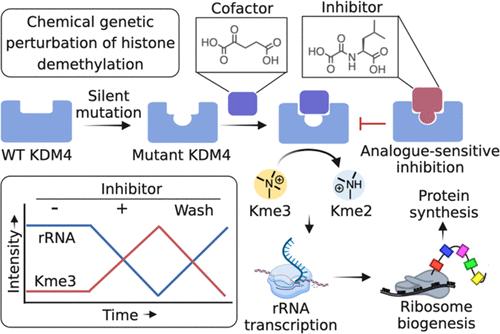
组蛋白去甲基化酶的类似敏感性抑制揭示了核糖体蛋白合成中的成员特异性功能
赖氨酸去甲基化酶(KDMs)利用地球上丰富的铁和代谢产物 2-氧代戊二酸(2OG)催化组蛋白中甲基的氧化脱除。KDMs 已成为真核基因表达的主调控因子和新的药物靶点;KDMs 的小分子抑制剂已进入治疗人类癌症的临床阶段。然而,对人类 KDMs 功能异质性的机理研究还很有限,因此有必要开发用于精准靶向的化学探针。在这里,我们鉴定了 KDM4 亚家族的类似物敏感(as)突变体,以时间定义的方式阐明成员的特异性生物功能。通过用丙氨酸取代 KDM4 亚家族成员活性位点上高度保守的苯丙氨酸残基,我们开发出了具有完整催化活性和底物特异性的突变体,其催化活性和底物特异性与野生型同源物无异。与野生型去甲基化酶不同,突变体对带有互补立体附属物的辅因子竞争性 N-草酰甘氨酸(NOG)类似物敏感。特别值得注意的是 N-草酰亮氨酸(NOL),它能以亚摩尔级的效力可逆地抑制 KDM4 突变体。细胞渗透性 NOL 原药可抑制培养人体细胞中的酶,从而调节核糖体组蛋白上的赖氨酸甲基化。通过对正交酶的条件性扰动,我们发现了 KDM4A 在核糖体蛋白质合成中的特异性作用,并绘制了一个涉及基因座特异性组蛋白去甲基化的显著动态信号级联,从而导致 rRNA 的快速表达、核糖体组装的增强以及蛋白质的合成。这些结果为 KDM4A 在癌症中的作用提供了机理线索,癌症依赖于核糖体活性的增强来支持不受控制的细胞增殖。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
CiteScore
24.40
自引率
6.00%
发文量
2398
审稿时长
1.6 months
期刊介绍:
The flagship journal of the American Chemical Society, known as the Journal of the American Chemical Society (JACS), has been a prestigious publication since its establishment in 1879. It holds a preeminent position in the field of chemistry and related interdisciplinary sciences. JACS is committed to disseminating cutting-edge research papers, covering a wide range of topics, and encompasses approximately 19,000 pages of Articles, Communications, and Perspectives annually. With a weekly publication frequency, JACS plays a vital role in advancing the field of chemistry by providing essential research.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


