Dynamic control of RNA-DNA hybrid formation orchestrates DNA2 activation at stalled forks by RNAPII and DDX39A
IF 16.6
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
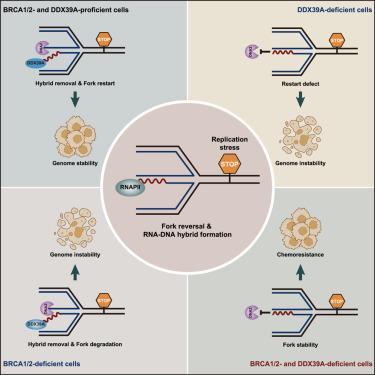
Stalled replication forks, susceptible to nucleolytic threats, necessitate protective mechanisms involving pivotal factors such as the tumor suppressors BRCA1 and BRCA2. Here, we demonstrate that, upon replication stress, RNA polymerase II (RNAPII) is recruited to stalled forks, actively promoting the transient formation of RNA-DNA hybrids. These hybrids act as safeguards, preventing premature engagement by the DNA2 nuclease and uncontrolled DNA2-mediated degradation of nascent DNA. Furthermore, we provide evidence that DExD box polypeptide 39A (DDX39A), serving as an RNA-DNA resolver, unwinds these structures and facilitates regulated DNA2 access to stalled forks. This orchestrated process enables controlled DNA2-dependent stalled fork processing and restart. Finally, we reveal that loss of DDX39A enhances stalled fork protection in BRCA1/2-deficient cells, consequently conferring chemoresistance. Our results suggest that the dynamic regulation of RNA-DNA hybrid formation at stalled forks by RNAPII and DDX39A precisely governs the timing of DNA2 activation, contributing to stalled fork protection, processing, and restart, ultimately promoting genome stability.
RNA-DNA杂交形成的动态控制通过RNAPII和DDX39A协调停滞分叉处的DNA2激活
停滞的复制分叉容易受到核溶解的威胁,需要涉及关键因子(如肿瘤抑制因子BRCA1和BRCA2)的保护机制。在这里,我们证明,在复制胁迫下,RNA聚合酶II (RNAPII)被招募到停滞分叉,积极促进RNA- dna杂交的短暂形成。这些杂交体起到保护作用,防止DNA2核酸酶过早接触和DNA2介导的未控制的新生DNA降解。此外,我们提供的证据表明,DExD盒多肽39A (DDX39A)作为RNA-DNA分解器,解开这些结构,并促进受调节的DNA2进入停滞的分叉。这个精心安排的过程可以控制依赖于dna的停顿分叉处理和重新启动。最后,我们揭示了DDX39A的缺失增强了brca1 /2缺陷细胞中停滞叉的保护,从而产生了化学耐药。我们的研究结果表明,RNAPII和DDX39A对停滞分叉处RNA-DNA杂交形成的动态调控精确地控制了DNA2激活的时间,有助于停滞分叉的保护、加工和重启,最终促进基因组的稳定。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


