Probing the limits and capabilities of diffusion models for the anatomic editing of digital twins
IF 12.4
1区 医学
Q1 HEALTH CARE SCIENCES & SERVICES
引用次数: 0
Abstract
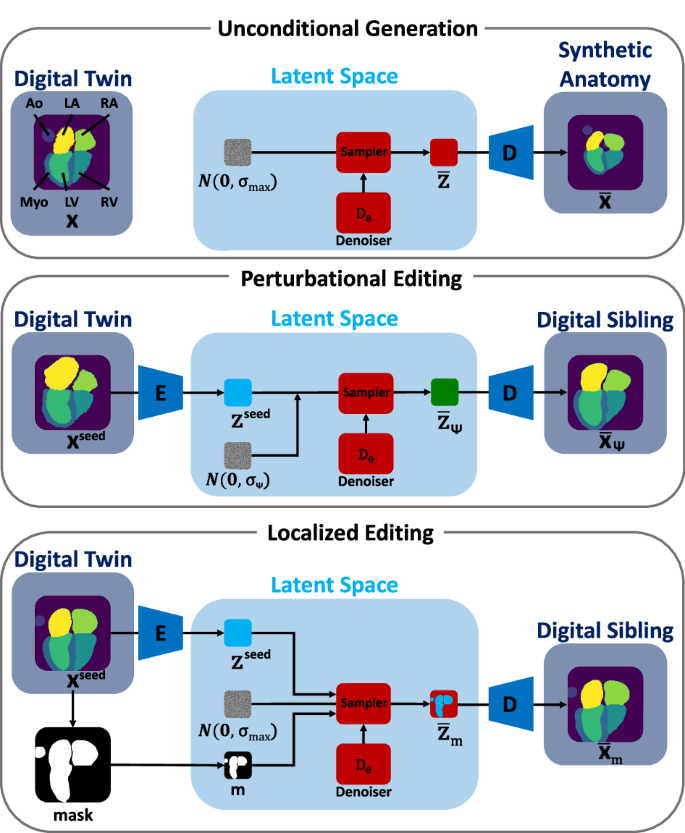
Numerical simulations of cardiovascular device deployment within digital twins of patient-specific anatomy can expedite and de-risk the device design process. Nonetheless, the exclusive use of patient-specific data constrains the anatomic variability that can be explored. We study how Latent Diffusion Models (LDMs) can edit digital twins to create digital siblings. Siblings can serve as the basis for comparative simulations, which can reveal how subtle anatomic variations impact device deployment, and augment virtual cohorts for improved device assessment. Using a case example centered on cardiac anatomy, we study various methods to generate digital siblings. We specifically introduce anatomic variation at different spatial scales or within localized regions, demonstrating the existence of bias toward common anatomic features. We furthermore leverage this bias for virtual cohort augmentation through selective editing, addressing issues related to dataset imbalance and diversity. Our framework delineates the capabilities of diffusion models in synthesizing anatomic variation for numerical simulation studies.

求助全文
约1分钟内获得全文
求助全文
来源期刊

NPJ Digital Medicine
Multiple-
CiteScore
25.10
自引率
3.30%
发文量
170
审稿时长
15 weeks
期刊介绍:
npj Digital Medicine is an online open-access journal that focuses on publishing peer-reviewed research in the field of digital medicine. The journal covers various aspects of digital medicine, including the application and implementation of digital and mobile technologies in clinical settings, virtual healthcare, and the use of artificial intelligence and informatics.
The primary goal of the journal is to support innovation and the advancement of healthcare through the integration of new digital and mobile technologies. When determining if a manuscript is suitable for publication, the journal considers four important criteria: novelty, clinical relevance, scientific rigor, and digital innovation.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


