Joining Natural and Synthetic DNA Using Biversal Nucleotides: Efficient Sequencing of Six-Nucleotide DNA
IF 14.4
1区 化学
Q1 CHEMISTRY, MULTIDISCIPLINARY
引用次数: 0
Abstract
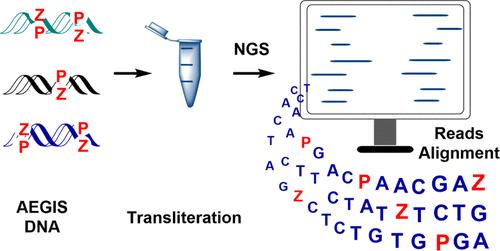
By rearranging hydrogen bond donor and acceptor groups within a standard Watson–Crick geometry, DNA can add eight independently replicable nucleotides forming four additional not found in standard Terran DNA. For many applications, the orthogonal pairing of standard and nonstandard pairs offers a key advantage. However, other applications require standard and nonstandard nucleotides to communicate with each other. This is especially true when seeking to recruit high-throughput instruments (e.g., Illumina), designed to sequence standard 4-nucleotide DNA, to sequence DNA that includes added nucleotides. For this purpose, PCR workflows are needed to replace nonstandard nucleotides in (for example) a 6-letter DNA sequence by defined mixtures of standard nucleotides built from 4 nucleotides. High-throughput sequencing can then report the sequences of those mixtures to bioinformatic alignment tools, which infer the original 6-nucleotide sequence by analysis of the mixtures. Unfortunately, the intrinsic orthogonality of standard and nonstandard nucleotides often demand polymerases that violate pairing biophysics to do this replacement, leading to inefficiencies in this “transliteration” process. Thus, laboratory in vitro evolution (LIVE) using “anthropogenic evolvable genetic information systems” (AEGIS), an important “consumer” of new sequencing tools, has been slow to be democratized; robust sequencing is needed to identify the AegisBodies and AegisZymes that AEGIS-LIVE delivers. This work introduces a new way to connect synthetic and standard molecular biology: biversal nucleotides. In an example presented here, a pyrimidine analogue (pyridine-2-one, y) pairs with Watson–Crick geometry to both a nonstandard base (2-amino-8-imidazo-[1,2a]-1,3,5-triazin-[8H]-4-one, P, the Watson–Crick partner of 6-amino-5-nitro-[1H]-pyridin-2-one, Z) and a base that completes the Watson–Crick hydrogen bond pattern (2-amino-2′-deoxyadenosine, amA). PCR amplification of GACTZP DNA with dyTP delivers products where Z:P pairs are cleanly transliterated to A:T pairs. In parallel, PCR of the same GACTZP sample at higher pH delivers products where Z:P pairs are cleanly transliterated to C:G pairs. By allowing robust sequencing of 6-letter GACTZP DNA, this workflow will help democratize AEGIS-LIVE. Further, other implementations of the biversal concept can enable communication across and between standard DNA and synthetic DNA more generally.
求助全文
约1分钟内获得全文
求助全文
来源期刊
CiteScore
24.40
自引率
6.00%
发文量
2398
审稿时长
1.6 months
期刊介绍:
The flagship journal of the American Chemical Society, known as the Journal of the American Chemical Society (JACS), has been a prestigious publication since its establishment in 1879. It holds a preeminent position in the field of chemistry and related interdisciplinary sciences. JACS is committed to disseminating cutting-edge research papers, covering a wide range of topics, and encompasses approximately 19,000 pages of Articles, Communications, and Perspectives annually. With a weekly publication frequency, JACS plays a vital role in advancing the field of chemistry by providing essential research.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


