Harnessing 12-lead ECG and MRI data to personalise repolarisation profiles in cardiac digital twin models for enhanced virtual drug testing
IF 10.7
1区 医学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
Abstract
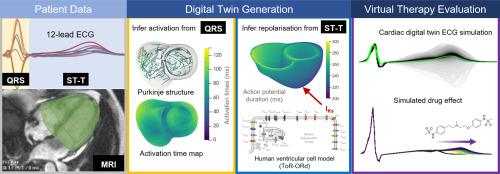
Cardiac digital twins are computational tools capturing key functional and anatomical characteristics of patient hearts for investigating disease phenotypes and predicting responses to therapy. When paired with large-scale computational resources and large clinical datasets, digital twin technology can enable virtual clinical trials on virtual cohorts to fast-track therapy development. Here, we present an open-source automated pipeline for personalising ventricular electrophysiological function based on routinely acquired magnetic resonance imaging (MRI) data and the standard 12-lead electrocardiogram (ECG).
Using MRI-based anatomical models, a sequential Monte-Carlo approximate Bayesian computational inference method is extended to infer electrical activation and repolarisation characteristics from the ECG. Fast simulations are conducted with a reaction-Eikonal model, including the Purkinje network and biophysically-detailed subcellular ionic current dynamics for repolarisation. For each patient, parameter uncertainty is represented by inferring an envelope of plausible ventricular models rather than a single one, which means that parameter uncertainty can be propagated to therapy evaluation. Furthermore, we have developed techniques for translating from reaction-Eikonal to monodomain simulations, which allows more realistic simulations of cardiac electrophysiology. The pipeline is demonstrated in three healthy subjects, where our inferred pseudo-diffusion reaction-Eikonal models reproduced the patient's ECG with a median Pearson's correlation coefficient of 0.9, and then translated to monodomain simulations with a median correlation coefficient of 0.84 across all subjects. We then demonstrate our digital twins for virtual evaluation of Dofetilide with uncertainty quantification. These evaluations using our cardiac digital twins reproduced dose-dependent QTc and T peak to T end prolongations that are in keeping with large population drug response data.
The methodologies for cardiac digital twinning presented here are a step towards personalised virtual therapy testing and can be scaled to generate virtual populations for clinical trials to fast-track therapy evaluation. The tools developed for this paper are open-source, documented, and made publicly available.
利用12导联心电图和MRI数据在心脏数字双胞胎模型中个性化复极谱,以增强虚拟药物测试
心脏数字双胞胎是一种计算工具,可以捕获患者心脏的关键功能和解剖特征,用于研究疾病表型和预测对治疗的反应。当与大规模计算资源和大型临床数据集相结合时,数字孪生技术可以使虚拟队列的虚拟临床试验快速跟踪治疗开发。在这里,我们提出了一个开源的自动化管道,用于个性化心室电生理功能,基于常规获得的磁共振成像(MRI)数据和标准12导联心电图(ECG)。利用基于mri的解剖模型,扩展了顺序蒙特卡罗近似贝叶斯计算推理方法,以推断ECG的电激活和复极特征。使用反应- eikonal模型进行快速模拟,包括浦肯野网络和重极化的生物物理详细亚细胞离子电流动力学。对于每个患者,参数不确定性是通过推断可行的心室模型的包络来表示的,而不是单一的,这意味着参数不确定性可以传播到治疗评估中。此外,我们已经开发了从反应- eikonal转换到单域模拟的技术,这允许更真实的心脏电生理模拟。该管道在三个健康受试者中得到了证明,我们推断的伪扩散反应- eikonal模型再现了患者的心电图,Pearson相关系数中值为0.9,然后在所有受试者中转化为单域模拟,相关系数中值为0.84。然后,我们展示了我们的数字双胞胎对不确定度量化的多非利特的虚拟评估。这些评估使用我们的心脏数字双胞胎再现了剂量依赖性QTc和T峰到T端延长,与大群体药物反应数据保持一致。这里介绍的心脏数字配对方法是向个性化虚拟治疗测试迈出的一步,可以扩展到为临床试验生成虚拟人群,以快速跟踪治疗评估。为本文开发的工具是开源的,有文档记录的,并且是公开可用的。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Medical image analysis
工程技术-工程:生物医学
CiteScore
22.10
自引率
6.40%
发文量
309
审稿时长
6.6 months
期刊介绍:
Medical Image Analysis serves as a platform for sharing new research findings in the realm of medical and biological image analysis, with a focus on applications of computer vision, virtual reality, and robotics to biomedical imaging challenges. The journal prioritizes the publication of high-quality, original papers contributing to the fundamental science of processing, analyzing, and utilizing medical and biological images. It welcomes approaches utilizing biomedical image datasets across all spatial scales, from molecular/cellular imaging to tissue/organ imaging.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


