Comparative transcriptomic analysis uncovers molecular heterogeneity in hepatobiliary cancers
IF 5
2区 医学
Q2 Medicine
引用次数: 0
Abstract
Hepatobiliary cancers (HBCs) pose a major global health challenge, with a lack of effective targeted biomarkers. Due to their complex anatomical locations, shared risk factors, and the limitations of targeted therapies, generalized treatment strategies are often used for gallbladder cancer (GBC), hepatocellular carcinoma (HCC), and intrahepatic cholangiocarcinoma (ICC). This study aimed to identify specific transcriptomic signatures in GBC, HCC, and ICC. The transcriptomic data analysis revealed distinct expression profiles, highlighting complex molecular heterogeneity within these cancers, even within the same organ system. Functional annotation revealed distinct biological pathways associated with each type of HBCs. GBC was linked to cell cycle regulation, HCC was associated with immune system modulation, and ICC was involved in metabolic dysregulation, particularly lipid metabolism. Gene co-expression network (GCN) and protein-protein interaction (PPI) network analyses identified potential key genes, such as MAPK3 and ERBB2 in GBC, AC069287.1 and ACTN2 in HCC, and TRPC1 and BACE1 in ICC. The FOX family of transcription factors (TFs) was conserved across all three cancer types. To further explore the relationship between Epithelial-Mesenchymal Transition (EMT) and the identified hub genes and TFs, an EMT score analysis was conducted. This analysis revealed distinct phenotypic characteristics in each cancer type, with TFs identified in GBC and ICC showing a stronger correlation with EMT compared to those in HCC. External validation using The Cancer Genome Atlas (TCGA) databases confirmed the expression of candidate genes, underscoring their potential as therapeutic targets. These findings provide valuable insights into the molecular heterogeneity and complexity of HBCs, opening new avenues for personalized therapeutic interventions.
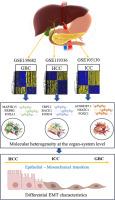
比较转录组分析揭示了肝胆癌的分子异质性。
由于缺乏有效的靶向生物标志物,肝胆癌(HBC)对全球健康构成了重大挑战。由于胆囊癌(GBC)、肝细胞癌(HCC)和肝内胆管癌(ICC)的解剖位置复杂、具有共同的风险因素以及靶向治疗的局限性,因此通常采用通用的治疗策略。本研究旨在确定 GBC、HCC 和 ICC 的特定转录组特征。转录组数据分析揭示了不同的表达谱,凸显了这些癌症内部复杂的分子异质性,即使在同一器官系统中也是如此。功能注释揭示了与每种类型的 HBC 相关的不同生物通路。GBC与细胞周期调控有关,HCC与免疫系统调节有关,而ICC则涉及代谢失调,尤其是脂质代谢。基因共表达网络(GCN)和蛋白相互作用(PPI)网络分析发现了潜在的关键基因,如GBC中的MAPK3和ERBB2、HCC中的AC069287.1和ACTN2以及ICC中的TRPC1和BACE1。FOX转录因子家族在所有三种癌症类型中都是保守的。为了进一步探讨上皮-间质转化(EMT)与已确定的枢纽基因和转录因子之间的关系,我们进行了一项EMT评分分析。该分析揭示了每种癌症类型的不同表型特征,与HCC相比,在GBC和ICC中发现的TF与EMT的相关性更强。利用癌症基因组图谱(TCGA)数据库进行的外部验证证实了候选基因的表达,凸显了它们作为治疗靶点的潜力。这些发现为了解 HBCs 的分子异质性和复杂性提供了有价值的见解,为个性化治疗干预开辟了新途径。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Translational Oncology
ONCOLOGY-
CiteScore
8.40
自引率
2.00%
发文量
314
审稿时长
54 days
期刊介绍:
Translational Oncology publishes the results of novel research investigations which bridge the laboratory and clinical settings including risk assessment, cellular and molecular characterization, prevention, detection, diagnosis and treatment of human cancers with the overall goal of improving the clinical care of oncology patients. Translational Oncology will publish laboratory studies of novel therapeutic interventions as well as clinical trials which evaluate new treatment paradigms for cancer. Peer reviewed manuscript types include Original Reports, Reviews and Editorials.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


