Enhancing chest X-ray datasets with privacy-preserving large language models and multi-type annotations: A data-driven approach for improved classification
IF 10.7
1区 医学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
Abstract
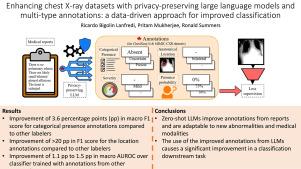
In chest X-ray (CXR) image analysis, rule-based systems are usually employed to extract labels from reports for dataset releases. However, there is still room for improvement in label quality. These labelers typically output only presence labels, sometimes with binary uncertainty indicators, which limits their usefulness. Supervised deep learning models have also been developed for report labeling but lack adaptability, similar to rule-based systems. In this work, we present MAPLEZ (Medical report Annotations with Privacy-preserving Large language model using Expeditious Zero shot answers), a novel approach leveraging a locally executable Large Language Model (LLM) to extract and enhance findings labels on CXR reports. MAPLEZ extracts not only binary labels indicating the presence or absence of a finding but also the location, severity, and radiologists’ uncertainty about the finding. Over eight abnormalities from five test sets, we show that our method can extract these annotations with an increase of 3.6 percentage points (pp) in macro F1 score for categorical presence annotations and more than 20 pp increase in F1 score for the location annotations over competing labelers. Additionally, using the combination of improved annotations and multi-type annotations in classification supervision in a dataset of limited-resolution CXRs, we demonstrate substantial advancements in proof-of-concept classification quality, with an increase of 1.1 pp in AUROC over models trained with annotations from the best alternative approach. We share code and annotations.
利用保护隐私的大型语言模型和多类型注释增强胸部 X 光数据集:改进分类的数据驱动方法。
在胸部 X 光(CXR)图像分析中,通常采用基于规则的系统从报告中提取标签,以便发布数据集。然而,标签质量仍有待提高。这些标签器通常只输出存在标签,有时还带有二进制不确定性指标,这限制了它们的实用性。也有人开发了用于报告标注的有监督深度学习模型,但与基于规则的系统类似,缺乏适应性。在这项工作中,我们提出了 MAPLEZ(使用快速零枪答案的隐私保护大语言模型医学报告注释),这是一种利用本地可执行大语言模型(LLM)来提取和增强 CXR 报告中的发现标签的新方法。MAPLEZ 不仅能提取二进制标签来表示有无发现,还能提取位置、严重程度和放射医师对发现的不确定性。在五个测试集中的八种异常情况中,我们证明了我们的方法可以提取这些注释,与竞争标签相比,分类存在注释的宏观 F1 分数提高了 3.6 个百分点,位置注释的 F1 分数提高了 20 多个百分点。此外,在有限分辨率 CXR 数据集的分类监督中结合使用改进注释和多类型注释,我们展示了概念验证分类质量的实质性进步,与使用最佳替代方法注释训练的模型相比,AUROC 提高了 1.1 个百分点。我们共享代码和注释。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Medical image analysis
工程技术-工程:生物医学
CiteScore
22.10
自引率
6.40%
发文量
309
审稿时长
6.6 months
期刊介绍:
Medical Image Analysis serves as a platform for sharing new research findings in the realm of medical and biological image analysis, with a focus on applications of computer vision, virtual reality, and robotics to biomedical imaging challenges. The journal prioritizes the publication of high-quality, original papers contributing to the fundamental science of processing, analyzing, and utilizing medical and biological images. It welcomes approaches utilizing biomedical image datasets across all spatial scales, from molecular/cellular imaging to tissue/organ imaging.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


