Structural basis for the effects of Ser387 phosphorylation of MgcRacGAP on its GTPase-activating activities for CDC42 and RHOA
IF 2.7
3区 生物学
Q3 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
MgcRacGAP is a GTPase-activating protein (GAP) for the Rho family GTPases. During cytokinesis, MgcRacGAP localizes to the midbody, where it activates the GTPase activity of Rho family GTPases to facilitate cytokinesis. In the midbody, Aurora B phosphorylates Ser387 within the GAP domain of human MgcRacGAP, a modification that is suggested to influence GTPase preference. However, there are conflicting reports, with some studies indicating that Ser387 phosphorylation does not alter the GTPase preference of MgcRacGAP. This controversy highlights the need for a deeper understanding of the molecular interactions involved, which can be clarified through structural analyses. In the present study, we determined the crystal structures of the wild-type MgcRacGAP GAP domain complexed with CDC42•GDP•AlF4− and the S378D phosphomimetic mutant GAP domain fused with RHOA•GDP•AlF4−. Additionally, crystal structures of the GAP domains were determined for the S387D and S387A mutants. Our analysis revealed that neither GTPase binding nor S387D mutation affected the overall structure of the GAP domain. However, comparison of the CDC42•MgcRacGAP (wild-type) complex with the RHOA-MgcRacGAP(S378D) fusion protein structure indicated that the S387D mutation caused positional shifts in both CDC42 and RHOA relative to MgcRacGAP. These shifts reduced interactions with CDC42 more severely than those with RHOA. In fact, the S387D mutation decreased the GTPase-activating activity of MgcRacGAP toward CDC42, while its impact on RHOA was only moderate. This difference in the rate of activity reduction may play an important role in GTPase preference.
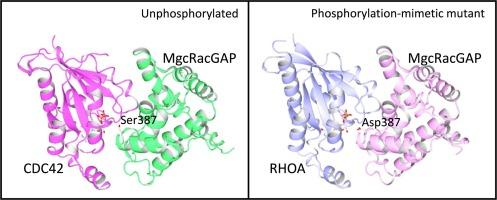
MgcRacGAP 的 Ser387 磷酸化对其 CDC42 和 RHOA 的 GTPase 激活活性的影响的结构基础。
MgcRacGAP是Rho家族GTP酶的一种GTP酶激活蛋白(GAP)。在细胞分裂过程中,MgcRacGAP 定位于中体,在那里激活 Rho 家族 GTPases 的 GTPase 活性,从而促进细胞分裂。在中体,极光 B 会磷酸化人 MgcRacGAP GAP 结构域内的 Ser387,这种修饰被认为会影响 GTPase 的偏好。然而,有一些研究表明,Ser387 磷酸化并不会改变 MgcRacGAP 的 GTPase 偏好。这种争议凸显了深入了解相关分子相互作用的必要性,而这可以通过结构分析来澄清。在本研究中,我们测定了野生型 MgcRacGAP GAP 结构域与 CDC42-GDP-AlF4- 复合物的晶体结构,以及 S378D 磷酸拟态突变体 GAP 结构域与 RHOA-GDP-AlF4- 融合的晶体结构。此外,我们还测定了 S387D 和 S387A 突变体 GAP 结构域的晶体结构。我们的分析表明,GTPase 结合和 S387D 突变都不会影响 GAP 结构域的整体结构。然而,CDC42-MgcRacGAP(野生型)复合物与 RHOA-MgcRacGAP(S378D)融合蛋白结构的比较表明,S387D 突变引起了 CDC42 和 RHOA 相对于 MgcRacGAP 的位置移动。与 RHOA 相比,这些位移更严重地降低了与 CDC42 的相互作用。事实上,S387D 突变降低了 MgcRacGAP 对 CDC42 的 GTPase 活化活性,而它对 RHOA 的影响只是温和的。这种活性降低速度上的差异可能在 GTPase 偏好中起着重要作用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of structural biology
生物-生化与分子生物学
CiteScore
6.30
自引率
3.30%
发文量
88
审稿时长
65 days
期刊介绍:
Journal of Structural Biology (JSB) has an open access mirror journal, the Journal of Structural Biology: X (JSBX), sharing the same aims and scope, editorial team, submission system and rigorous peer review. Since both journals share the same editorial system, you may submit your manuscript via either journal homepage. You will be prompted during submission (and revision) to choose in which to publish your article. The editors and reviewers are not aware of the choice you made until the article has been published online. JSB and JSBX publish papers dealing with the structural analysis of living material at every level of organization by all methods that lead to an understanding of biological function in terms of molecular and supermolecular structure.
Techniques covered include:
• Light microscopy including confocal microscopy
• All types of electron microscopy
• X-ray diffraction
• Nuclear magnetic resonance
• Scanning force microscopy, scanning probe microscopy, and tunneling microscopy
• Digital image processing
• Computational insights into structure
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


