Transcriptome analysis uncovers the expression of genes associated with growth in the gills and muscles of white shrimp (Litopenaeus vannamei) with different growth rates
IF 2.2
2区 生物学
Q4 BIOCHEMISTRY & MOLECULAR BIOLOGY
Comparative Biochemistry and Physiology D-Genomics & Proteomics
Pub Date : 2024-10-28
DOI:10.1016/j.cbd.2024.101347
引用次数: 0
Abstract
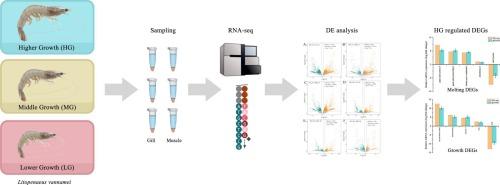
Litopenaeus vannamei is a crucial species in aquaculture. The gene expression patterns associated with distinct growth rates are not well understood. To investigate this, we used RNA-seq to study the underlying growth mechanism of L. vannamei with varying growth rates. Individuals of higher growth performance (HG), middle growth performance (MG), and lower growth performance (LG) were examined. A total of 8422 and 4560 differentially expressed genes (DEGs) were identified in gill and muscle samples, respectively. Genes related to growth were significantly up-regulated in HG gills, such as cuticle protein, chitin synthase, pupal cuticle protein, titin myosin G heavy chain, and myosin heavy chain 10. The GO enrichment analysis revealed that the DEGs of HG gills were significantly enriched in “structural constituent of cuticle”, “primary metabolic process” and “chitin binding”. The growth-related genes were highly expressed in HG muscle, such as myosin heavy chain, myosin heavy chain type A and myosin 3. The GO enrichment analysis revealed that the DEGs of HG muscle were significantly enriched in “myosin filament”, “myosin complex” and “myofibril”. These findings provide insights into mechanisms underlying the growth performance of superior L. vannamei, and identify candidate genes for genetic improvement programs aimed at enhancing this trait.
转录组分析揭示了不同生长速度的南美白对虾(Litopenaeus vannamei)鳃和肌肉中与生长相关的基因的表达情况
凡纳滨对虾是水产养殖中的重要物种。与不同生长率相关的基因表达模式尚不十分清楚。为了研究这个问题,我们使用 RNA-seq 研究了不同生长率的凡纳滨对虾的潜在生长机制。我们研究了生长性能较高(HG)、生长性能中等(MG)和生长性能较低(LG)的个体。在鳃和肌肉样本中分别发现了 8422 个和 4560 个差异表达基因(DEG)。与生长相关的基因在HG鳃中明显上调,如角质层蛋白、几丁质合成酶、蛹角质层蛋白、titin肌球蛋白G重链和肌球蛋白重链10。GO富集分析表明,HG鳃的DEGs在 "角质层结构成分"、"初级代谢过程 "和 "几丁质结合 "中显著富集。GO富集分析表明,HG肌肉中的DEGs在 "肌球蛋白丝"、"肌球蛋白复合物 "和 "肌原纤维 "中明显富集。这些发现有助于深入了解优质凡纳米鱼生长性能的内在机制,并为旨在提高该性状的遗传改良计划确定候选基因。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
CiteScore
5.10
自引率
3.30%
发文量
69
审稿时长
33 days
期刊介绍:
Comparative Biochemistry & Physiology (CBP) publishes papers in comparative, environmental and evolutionary physiology.
Part D: Genomics and Proteomics (CBPD), focuses on “omics” approaches to physiology, including comparative and functional genomics, metagenomics, transcriptomics, proteomics, metabolomics, and lipidomics. Most studies employ “omics” and/or system biology to test specific hypotheses about molecular and biochemical mechanisms underlying physiological responses to the environment. We encourage papers that address fundamental questions in comparative physiology and biochemistry rather than studies with a focus that is purely technical, methodological or descriptive in nature.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


