COLD6-OSM1 module senses chilling for cold tolerance via 2′,3′-cAMP signaling in rice
IF 14.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
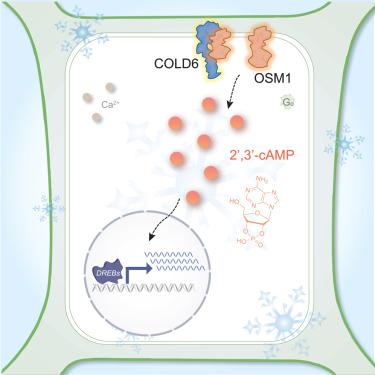
While it is known that temperature sensors trigger calcium (Ca2+) signaling to confer cold tolerance in cells, less is known about sensors that couple with other secondary messengers. Here, we identify a cold sensor complex of CHILLING-TOLERANCE DIVERGENCE 6 (COLD6) and osmotin-like 1 (OSM1), which triggers 2′,3′-cyclic adenosine monophosphate (2′,3′-cAMP) production to enhance cold tolerance in rice. COLD6, which is encoded by a major quantitative trait locus (QTL) gene, interacts with the rice G protein α subunit (RGA1) at the plasma membrane under normal conditions. Upon exposure to chilling, cold-induced OSM1 binds to COLD6, kicking out RGA1 from interaction. This triggers an elevation of 2′,3′-cAMP levels for enhancing chilling tolerance. Genetic data show that COLD6 negatively regulates cold tolerance and functionally depends on OSM1 in chilling stress. COLD6 alleles were selected during rice domestication. Knockout and natural variation of COLD6 in hybrid rice enhanced chilling tolerance, hinting design potential for breeding. This highlighted a module triggering 2′,3′-cAMP to improve chilling tolerance in crops.
水稻中的 COLD6-OSM1 模块通过 2′,3′-cAMP 信号传导感知寒冷以提高耐寒性
众所周知,温度传感器会触发钙(Ca2+)信号传导,从而赋予细胞耐寒性,但人们对与其他次级信使耦合的传感器却知之甚少。在这里,我们发现了一个由寒冷-耐受分化6(COLD6)和类渗透蛋白1(OSM1)组成的冷传感器复合物,它能触发2′,3′-环磷酸腺苷(2′,3′-cAMP)的产生,从而增强水稻的耐寒性。COLD6 由一个主要的数量性状基因座(QTL)基因编码,在正常情况下与质膜上的水稻 G 蛋白 α 亚基(RGA1)相互作用。在寒冷条件下,冷诱导的 OSM1 与 COLD6 结合,将 RGA1 从相互作用中踢出。这引发了 2′,3′-cAMP 水平的升高,从而增强了耐寒性。遗传数据表明,COLD6 负向调节耐寒性,并在寒冷胁迫中对 OSM1 起作用。COLD6 等位基因是在水稻驯化过程中筛选出来的。杂交水稻中 COLD6 的敲除和自然变异增强了耐寒性,为育种设计提供了潜力。这突显了一个触发 2′,3′-cAMP 的模块可提高作物的耐寒性。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Molecular Cell
生物-生化与分子生物学
CiteScore
26.00
自引率
3.80%
发文量
389
审稿时长
1 months
期刊介绍:
Molecular Cell is a companion to Cell, the leading journal of biology and the highest-impact journal in the world. Launched in December 1997 and published monthly. Molecular Cell is dedicated to publishing cutting-edge research in molecular biology, focusing on fundamental cellular processes. The journal encompasses a wide range of topics, including DNA replication, recombination, and repair; Chromatin biology and genome organization; Transcription; RNA processing and decay; Non-coding RNA function; Translation; Protein folding, modification, and quality control; Signal transduction pathways; Cell cycle and checkpoints; Cell death; Autophagy; Metabolism.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


