Vitis vinifera Lipoxygenase LoxA is an Allosteric Dimer Activated by Lipidic Surfaces
IF 4.7
2区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Lipoxygenases catalyze the peroxidation of poly-unsaturated fatty acid chains either free or esterified in membrane lipids. Vitis vinifera LoxA is transcriptionally induced at ripening onset and localizes at the inner chloroplast membrane where it is responsible for galactolipid regiospecific mono- and di-peroxidation. Here we present a kinetic and structural characterization of LoxA. Our X-ray structures reveal a constitutive dimer with detergent induced conformational changes affecting substrate binding and catalysis. In a closed conformation, a LID domain prevents substrate access to the catalytic site by steric hindrance. Detergent addition above the CMC destabilizes the LID and opens the dimer with both catalytic sites accessible from the same surface framed by the PLAT domains. As a consequence, detergent molecules occupy allosteric sites in the PLAT/catalytic domain interface. These structural changes are mirrored by increased enzymatic activity and positive cooperativity when the substrate is provided in micelles. The ability to interact with micelles is lost upon dimer destabilization by site-directed mutagenesis as assessed by tryptophan fluorescence. Our data allow to propose a model for protein activation at the membrane, classifying LoxA as an interfacial enzyme acting on fatty acid chains directly from the membrane similar to mammalian 15-LOX and 5-LOX.
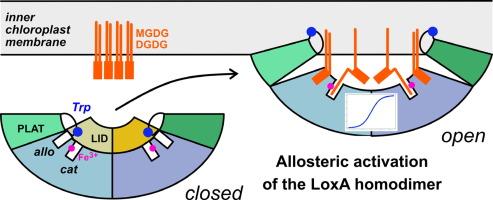
葡萄脂氧合酶 LoxA 是由脂质表面激活的异位二聚体。
脂氧合酶催化膜脂中游离或酯化的多不饱和脂肪酸链发生过氧化反应。葡萄 LoxA 在葡萄成熟时转录诱导,定位于叶绿体内膜,负责半乳糖脂特异性单过氧化和二过氧化。在这里,我们介绍了 LoxA 的动力学和结构特征。我们的 X 射线结构揭示了一个组成型二聚体,其构象在洗涤剂的诱导下发生了变化,从而影响了底物结合和催化作用。在封闭构象中,一个 LID 结构域通过立体阻碍阻止底物进入催化位点。洗涤剂添加到 CMC 以上会破坏 LID 的稳定性,并打开二聚体,使两个催化位点都能从 PLAT 结构域框定的同一表面进入。因此,洗涤剂分子占据了 PLAT/催化结构域界面上的异构位点。在胶束中提供底物时,酶活性和正合作性都会增加,从而反映出这些结构变化。通过色氨酸荧光评估,当定点突变使二聚体不稳定时,与胶束相互作用的能力就会丧失。根据我们的数据,可以提出一个膜上蛋白质活化模型,将 LoxA 归类为一种直接从膜上作用于脂肪酸链的界面酶,类似于哺乳动物的 15-LOX 和 5-LOX。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Journal of Molecular Biology
生物-生化与分子生物学
CiteScore
11.30
自引率
1.80%
发文量
412
审稿时长
28 days
期刊介绍:
Journal of Molecular Biology (JMB) provides high quality, comprehensive and broad coverage in all areas of molecular biology. The journal publishes original scientific research papers that provide mechanistic and functional insights and report a significant advance to the field. The journal encourages the submission of multidisciplinary studies that use complementary experimental and computational approaches to address challenging biological questions.
Research areas include but are not limited to: Biomolecular interactions, signaling networks, systems biology; Cell cycle, cell growth, cell differentiation; Cell death, autophagy; Cell signaling and regulation; Chemical biology; Computational biology, in combination with experimental studies; DNA replication, repair, and recombination; Development, regenerative biology, mechanistic and functional studies of stem cells; Epigenetics, chromatin structure and function; Gene expression; Membrane processes, cell surface proteins and cell-cell interactions; Methodological advances, both experimental and theoretical, including databases; Microbiology, virology, and interactions with the host or environment; Microbiota mechanistic and functional studies; Nuclear organization; Post-translational modifications, proteomics; Processing and function of biologically important macromolecules and complexes; Molecular basis of disease; RNA processing, structure and functions of non-coding RNAs, transcription; Sorting, spatiotemporal organization, trafficking; Structural biology; Synthetic biology; Translation, protein folding, chaperones, protein degradation and quality control.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


