A Hydrophobic Core Stabilizes the Residual Structure in the RRM2 Intermediate State of the ALS-linked Protein TDP-43
IF 4.7
2区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Folding intermediates mediate both protein folding and the misfolding and aggregation observed in human diseases, including amyotrophic lateral sclerosis (ALS), and are prime targets for therapeutic interventions. In this study, we identified the core nucleus of structure for a folding intermediate in the second RNA recognition motif (RRM2) of the ALS-linked RNA-binding protein, TDP-43 (TAR DNA-binding protein-43), using a combination of experimental and computational approaches. Urea equilibrium unfolding studies revealed that the RRM2 intermediate state consists of collapsed residual secondary structure localized to the N-terminal half of RRM2, while the C-terminus is largely disordered. Steered molecular dynamics simulations and mutagenesis studies yielded key stabilizing hydrophobic contacts that, when mutated to alanine, severely disrupt the overall fold of RRM2. In combination, these findings suggest a role for this RRM intermediate in normal TDP-43 function as well as serving as a template for misfolding and aggregation through the low stability and non-native secondary structure.
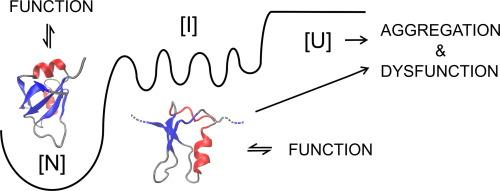
疏水核心稳定了 ALS 链接蛋白 TDP-43 的 RRM2 中间状态的残余结构。
折叠中间体既介导蛋白质折叠,也介导包括肌萎缩性脊髓侧索硬化症(ALS)在内的人类疾病中观察到的错误折叠和聚集,是治疗干预的主要目标。在这项研究中,我们采用实验和计算相结合的方法,在与 ALS 相关的 RNA 结合蛋白 TDP-43 的第二个 RNA 识别基序(RRM2)中确定了折叠中间体的核心结构核。尿素平衡展开研究发现,RRM2 中间状态由坍塌的残余二级结构组成,这些二级结构位于 RRM2 的 N 端半部分,而 C 端大部分处于无序状态。这些研究结果表明,RRM2 中间体在正常的 TDP-43 功能中发挥作用,并通过低稳定性和非原生二级结构成为错误折叠和聚集的模板。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Journal of Molecular Biology
生物-生化与分子生物学
CiteScore
11.30
自引率
1.80%
发文量
412
审稿时长
28 days
期刊介绍:
Journal of Molecular Biology (JMB) provides high quality, comprehensive and broad coverage in all areas of molecular biology. The journal publishes original scientific research papers that provide mechanistic and functional insights and report a significant advance to the field. The journal encourages the submission of multidisciplinary studies that use complementary experimental and computational approaches to address challenging biological questions.
Research areas include but are not limited to: Biomolecular interactions, signaling networks, systems biology; Cell cycle, cell growth, cell differentiation; Cell death, autophagy; Cell signaling and regulation; Chemical biology; Computational biology, in combination with experimental studies; DNA replication, repair, and recombination; Development, regenerative biology, mechanistic and functional studies of stem cells; Epigenetics, chromatin structure and function; Gene expression; Membrane processes, cell surface proteins and cell-cell interactions; Methodological advances, both experimental and theoretical, including databases; Microbiology, virology, and interactions with the host or environment; Microbiota mechanistic and functional studies; Nuclear organization; Post-translational modifications, proteomics; Processing and function of biologically important macromolecules and complexes; Molecular basis of disease; RNA processing, structure and functions of non-coding RNAs, transcription; Sorting, spatiotemporal organization, trafficking; Structural biology; Synthetic biology; Translation, protein folding, chaperones, protein degradation and quality control.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


