Genetics, epigenetics and autoimmunity constitute a Bermuda triangle for the pathogenesis of rheumatoid arthritis
IF 5.2
2区 医学
Q1 MEDICINE, RESEARCH & EXPERIMENTAL
引用次数: 0
Abstract
Rheumatoid arthritis (RA), a multigene disorder with a heritability rate of 60 %, is characterized by persistent pain, synovial hyperplasia, and cartilage and bone destruction, ultimately causing irreversible joint deformity. The etiology and pathogenesis of rheumatoid arthritis (RA) are primarily influenced by specific genetic variants, particularly HLA alleles such as HLA-DRB1*01 and DRB1*04. However, other HLA alleles such as HLA-DRB1*10 and DPB*1 have also been found to contribute to increased susceptibility to RA. However, non-HLA genes also confer a comparatively high risk of RA disease manifestation. The most relevant single nucleotide polymorphisms (SNPs) associated with non-HLA genes are PTPN22, TRAF1, CXCL-12, TBX-5, STAT4, FCGR, PADI4, and MTHFR. In conjunction with genetic susceptibility, epigenetic alterations orchestrate paramount involvement in regulating RA pathogenesis. Increasing evidence implicates DNA methylation and histone protein modifications, including acetylation and methylation, as the primary epigenetic mechanisms that drive the pathogenesis and clinical progression of the disease. In addition to genetic and epigenetic changes, autoimmune inflammation also determines the pathological progression of the synovial membrane in joints with RA. Glycosylation changes, such as sialylation and fucosylation, in immune cells have been shown to be relevant to disease progression. Genetic heterogeneity, epigenetic factors, and changes in glycosylation do not fully explain the features of RA. Therefore, investigating the interplay between genetics, epigenetics, and autoimmunity is crucial. This review highlights the significance and interaction of these elements in RA pathophysiology, suggesting their diagnostic potential and opening new avenues for novel therapeutic approaches.
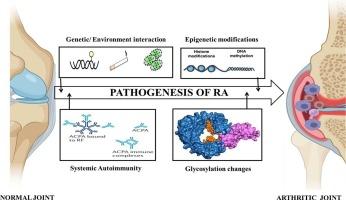
遗传学、表观遗传学和自身免疫学构成了类风湿关节炎发病机制的百慕大三角。
类风湿性关节炎(RA)是一种多基因疾病,遗传率高达 60%,以持续疼痛、滑膜增生、软骨和骨破坏为特征,最终导致不可逆转的关节畸形。类风湿性关节炎(RA)的病因和发病机制主要受特定遗传变异的影响,尤其是 HLA 等位基因,如 HLA-DRB1*01 和 DRB1*04。不过,也发现其他 HLA 等位基因,如 HLA-DRB1*10 和 DPB*1 也会导致 RA 易感性增加。然而,非 HLA 基因也会导致相对较高的 RA 发病风险。与非 HLA 基因最相关的单核苷酸多态性(SNPs)包括 PTPN22、TRAF1、CXCL-12、TBX-5、STAT4、FCGR、PADI4 和 MTHFR。除遗传易感性外,表观遗传学改变也是调节 RA 发病机制的重要因素。越来越多的证据表明,DNA甲基化和组蛋白修饰(包括乙酰化和甲基化)是驱动疾病发病和临床进展的主要表观遗传机制。除了遗传和表观遗传学变化外,自身免疫性炎症也决定了 RA 关节滑膜的病理进展。免疫细胞中的糖基化变化,如糖基化和岩藻糖基化,已被证明与疾病进展有关。遗传异质性、表观遗传因素和糖基化变化并不能完全解释 RA 的特征。因此,研究遗传学、表观遗传学和自身免疫学之间的相互作用至关重要。本综述强调了这些因素在RA病理生理学中的重要性和相互作用,提示了它们的诊断潜力,并为新型治疗方法开辟了新途径。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Life sciences
医学-药学
CiteScore
12.20
自引率
1.60%
发文量
841
审稿时长
6 months
期刊介绍:
Life Sciences is an international journal publishing articles that emphasize the molecular, cellular, and functional basis of therapy. The journal emphasizes the understanding of mechanism that is relevant to all aspects of human disease and translation to patients. All articles are rigorously reviewed.
The Journal favors publication of full-length papers where modern scientific technologies are used to explain molecular, cellular and physiological mechanisms. Articles that merely report observations are rarely accepted. Recommendations from the Declaration of Helsinki or NIH guidelines for care and use of laboratory animals must be adhered to. Articles should be written at a level accessible to readers who are non-specialists in the topic of the article themselves, but who are interested in the research. The Journal welcomes reviews on topics of wide interest to investigators in the life sciences. We particularly encourage submission of brief, focused reviews containing high-quality artwork and require the use of mechanistic summary diagrams.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


