π-π Interactions Drive the Homotypic Phase Separation of the Prion-like Diatom Pyrenoid Scaffold PYCO1
IF 4.7
2区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
CO2 fixation in most unicellular algae relies on the pyrenoid, a biomolecular condensate, which sequesters the cell’s carboxylase Rubisco. In the marine diatom Phaeodactylum tricornutum, the pyrenoid tandem repeat protein Pyrenoid Component 1 (PYCO1) multivalently binds Rubisco to form a heterotypic Rubisco condensate. PYCO1 contains prion-like domains and can phase-separate homotypically in a salt-dependent manner. Here we dissect PYCO1 homotypic liquid–liquid phase separation (LLPS) by evaluating protein fragments and the effect of site-directed mutagenesis. Two of PYCO1′s six repeats are required for homotypic LLPS. Mutagenesis of a minimal phase-separating fragment reveals tremendous sensitivity to the substitution of aromatic residues. Removing positively charged lysines and arginines instead enhances the propensity of the fragment to condense. We conclude that PYCO1 homotypic LLPS is mostly driven by π-π interactions mediated by tyrosine and tryptophan stickers. In contrast π-cation interactions involving arginine or lysine are not significant drivers of LLPS in this system.
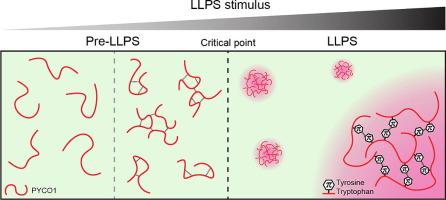
π-π相互作用驱动朊病毒样硅藻火绒素支架PYCO1的同型相分离。
大多数单细胞藻类的二氧化碳固定都依赖于类肾上腺素(一种生物分子凝结物),它能封存细胞的羧化酶 Rubisco。在海洋硅藻 Phaeodactylum tricornutum 中,类肾上腺素串联重复蛋白类肾上腺素成分 1(PYCO1)可多价结合 Rubisco,形成异型 Rubisco 凝聚物。PYCO1含有朊病毒样结构域,能以盐依赖的方式进行同型相分离。在这里,我们通过评估蛋白质片段和定点突变的效果来剖析PYCO1同型液-液相分离(LLPS)。同型液-液相分离需要PYCO1六个重复序列中的两个。对最小相分离片段的突变显示了对芳香族残基替代的极大敏感性。去除带正电荷的赖氨酸和精氨酸反而会增强该片段的凝聚倾向。我们的结论是,PYCO1 同型 LLPS 主要是由酪氨酸和色氨酸粘合剂介导的 π-π 相互作用驱动的。相比之下,涉及精氨酸或赖氨酸的π-阳离子相互作用在该系统中并不是 LLPS 的重要驱动力。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Journal of Molecular Biology
生物-生化与分子生物学
CiteScore
11.30
自引率
1.80%
发文量
412
审稿时长
28 days
期刊介绍:
Journal of Molecular Biology (JMB) provides high quality, comprehensive and broad coverage in all areas of molecular biology. The journal publishes original scientific research papers that provide mechanistic and functional insights and report a significant advance to the field. The journal encourages the submission of multidisciplinary studies that use complementary experimental and computational approaches to address challenging biological questions.
Research areas include but are not limited to: Biomolecular interactions, signaling networks, systems biology; Cell cycle, cell growth, cell differentiation; Cell death, autophagy; Cell signaling and regulation; Chemical biology; Computational biology, in combination with experimental studies; DNA replication, repair, and recombination; Development, regenerative biology, mechanistic and functional studies of stem cells; Epigenetics, chromatin structure and function; Gene expression; Membrane processes, cell surface proteins and cell-cell interactions; Methodological advances, both experimental and theoretical, including databases; Microbiology, virology, and interactions with the host or environment; Microbiota mechanistic and functional studies; Nuclear organization; Post-translational modifications, proteomics; Processing and function of biologically important macromolecules and complexes; Molecular basis of disease; RNA processing, structure and functions of non-coding RNAs, transcription; Sorting, spatiotemporal organization, trafficking; Structural biology; Synthetic biology; Translation, protein folding, chaperones, protein degradation and quality control.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


