New Catalytic Residues and Catalytic Mechanism of the RNase T1 Family
IF 3.8
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
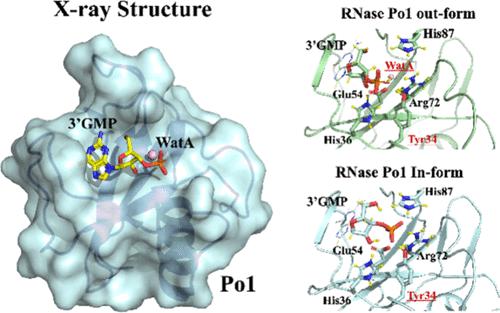
The ribonuclease T1 family, including RNase Po1 secreted by Pleurotus ostreatus, exhibits antitumor activity. Here, we resolved the Po1/guanosine-3′-monophosphate complex (3′GMP) structure at 1.75 Å. Structure comparison and fragment molecular orbital (FMO) calculation between the apo form and the Po1/3′GMP complex identified Phe38, Phe40, and Glu42 as the key binding residues. Two types of the RNase/3′GMP complex in RNasePo1 and RNase T1 were homologous to Po1, and FMO calculations elucidated that the biprotonated histidine on the β3 sheet (His36) on the β3 sheet and deprotonated Glu54 on the β4 sheet were advantageous to RNase activity. Moreover, tyrosine (Tyr34) on the β3 sheet was elucidated as a crucial catalytic residues. Mutation of Tyr34 with phenylalanine decreased RNase activity and diminished antitumor efficacy compared to that in the wild type. This suggests the importance of RNase activity in antitumor mechanisms.
RNase T1 家族的新催化残基和催化机理
核糖核酸酶 T1 家族(包括由梭梭菌分泌的 RNase Po1)具有抗肿瘤活性。通过结构比较和片段分子轨道(FMO)计算,我们确定了Phe38、Phe40和Glu42是Po1/鸟苷-3′-单磷酸复合物(3′GMP)的关键结合残基。RNasePo1 和 RNase T1 中两种类型的 RNase/3′GMP 复合物与 Po1 同源,FMO 计算表明,β3 片层上的双质子化组氨酸(His36)和β4 片层上的去质子化 Glu54 对 RNase 活性有利。此外,β3 片层上的酪氨酸(Tyr34)被认为是一个关键的催化残基。与野生型相比,用苯丙氨酸突变 Tyr34 会降低 RNase 的活性和抗肿瘤效果。这表明了 RNase 活性在抗肿瘤机制中的重要性。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

ACS Bio & Med Chem Au
药物、生物、化学-
CiteScore
4.10
自引率
0.00%
发文量
0
期刊介绍:
ACS Bio & Med Chem Au is a broad scope open access journal which publishes short letters comprehensive articles reviews and perspectives in all aspects of biological and medicinal chemistry. Studies providing fundamental insights or describing novel syntheses as well as clinical or other applications-based work are welcomed.This broad scope includes experimental and theoretical studies on the chemical physical mechanistic and/or structural basis of biological or cell function in all domains of life. It encompasses the fields of chemical biology synthetic biology disease biology cell biology agriculture and food natural products research nucleic acid biology neuroscience structural biology and biophysics.The journal publishes studies that pertain to a broad range of medicinal chemistry including compound design and optimization biological evaluation molecular mechanistic understanding of drug delivery and drug delivery systems imaging agents and pharmacology and translational science of both small and large bioactive molecules. Novel computational cheminformatics and structural studies for the identification (or structure-activity relationship analysis) of bioactive molecules ligands and their targets are also welcome. The journal will consider computational studies applying established computational methods but only in combination with novel and original experimental data (e.g. in cases where new compounds have been designed and tested).Also included in the scope of the journal are articles relating to infectious diseases research on pathogens host-pathogen interactions therapeutics diagnostics vaccines drug-delivery systems and other biomedical technology development pertaining to infectious diseases.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


