Automated customization of large-scale spiking network models to neuronal population activity
IF 18.3
Q1 COMPUTER SCIENCE, INTERDISCIPLINARY APPLICATIONS
引用次数: 0
Abstract
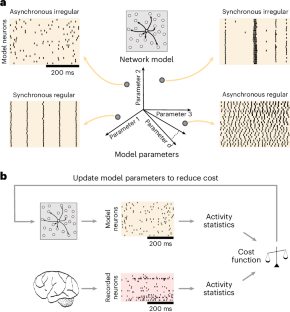
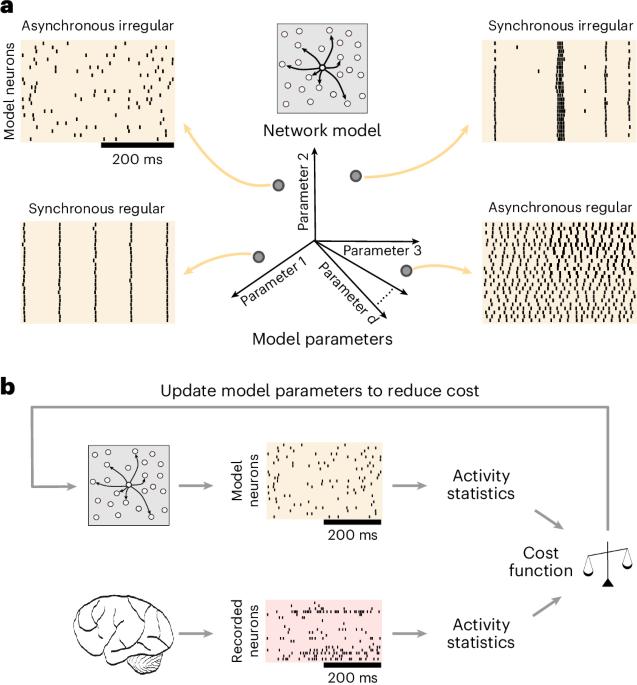
Understanding brain function is facilitated by constructing computational models that accurately reproduce aspects of brain activity. Networks of spiking neurons capture the underlying biophysics of neuronal circuits, yet their activity’s dependence on model parameters is notoriously complex. As a result, heuristic methods have been used to configure spiking network models, which can lead to an inability to discover activity regimes complex enough to match large-scale neuronal recordings. Here we propose an automatic procedure, Spiking Network Optimization using Population Statistics (SNOPS), to customize spiking network models that reproduce the population-wide covariability of large-scale neuronal recordings. We first confirmed that SNOPS accurately recovers simulated neural activity statistics. Then, we applied SNOPS to recordings in macaque visual and prefrontal cortices and discovered previously unknown limitations of spiking network models. Taken together, SNOPS can guide the development of network models, thereby enabling deeper insight into how networks of neurons give rise to brain function. An automatic framework, SNOPS, is developed for configuring a spiking network model to reproduce neuronal recordings. It is used to discover previously unknown limitations of spiking network models, thereby guiding model development.
根据神经元群体活动自动定制大规模尖峰网络模型
构建能准确再现大脑活动的计算模型有助于理解大脑功能。尖峰神经元网络捕捉到了神经元回路的基本生物物理学原理,但它们的活动对模型参数的依赖却出了名的复杂。因此,启发式方法一直被用于配置尖峰网络模型,这可能导致无法发现足够复杂的活动机制,从而无法与大规模神经元记录相匹配。在这里,我们提出了一种自动程序--使用群体统计的尖峰网络优化(SNOPS)--来定制尖峰网络模型,以重现大规模神经元记录的群体共变性。我们首先证实 SNOPS 能准确恢复模拟的神经活动统计数据。然后,我们将 SNOPS 应用于猕猴视觉和前额叶皮层的记录,发现了尖峰网络模型之前未知的局限性。综上所述,SNOPS 可以指导网络模型的开发,从而让人们更深入地了解神经元网络是如何产生大脑功能的。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


