{"title":"A Cost-Effective Approach for Single-Stranded DNA Amplification Using Primer-Blocked Asymmetric PCR","authors":"Krisztina Percze, Ákos Harkai, Tamás Mészáros","doi":"10.1002/cpz1.1125","DOIUrl":null,"url":null,"abstract":"<p>In vitro amplification of single-stranded oligonucleotide libraries presents a significant challenge due to the potential for excessive byproduct formation. This phenomenon largely affects the quality of the ssDNAs created using the most commonly used methods, e.g., asymmetric PCR, biotin-streptavidin separation, or lambda exonuclease digestion of dsDNA. Here, we describe an improved protocol that combines primer-blocked asymmetric PCR (PBA-PCR) with emulsion PCR and a cost-effective downstream process that altogether alleviates byproduct formation without distorting the sequence space of the ssDNA library. In PBA-PCR, the reaction mixture is complemented with a 3′-phosphate-blocked limiting primer that decreases mispriming, thus reducing polymerization of DNA byproducts. The downstream process includes mixing of the PBA-PCR product with excess reverse complement of the 3′-phosphate-blocked limiting primer and removal of dsDNA strands via biotin-streptavidin separation, yielding purified ssDNAs. In conclusion, we have devised a universally applicable approach for simple and cost-effective production of ssDNA libraries and unique ssDNA sequences with on-demand labeling. Our protocol could be beneficial for a variety of uses, such as generating aptamer libraries for SELEX, creating unique molecular identifiers for a wide range of sequencing applications, providing donor DNA for CRISPR-Cas9 systems, developing scaffold nanostructures, and enabling DNA-based data storage. © 2024 The Author(s). Current Protocols published by Wiley Periodicals LLC.</p><p><b>Basic Protocol 1</b>: Amplification of ssDNA libraries using PBA-PCR</p><p><b>Alternate Protocol 1</b>: Amplification of ssDNA libraries using emulsion PBA-PCR with a simplified extraction of PBA-PCR products</p><p><b>Basic Protocol 2</b>: Purification of PBA-PCR products to remove dsDNA and conversion of 3′-blocked primer to double-stranded complexes</p><p><b>Alternate Protocol 2</b>: Purification of PBA-PCR products to remove both dsDNA and blocking primers from the reaction mixture</p><p><b>Support Protocol</b>: Analysis of PBA-PCR products by gel electrophoresis</p>","PeriodicalId":93970,"journal":{"name":"Current protocols","volume":"4 9","pages":""},"PeriodicalIF":0.0000,"publicationDate":"2024-09-04","publicationTypes":"Journal Article","fieldsOfStudy":null,"isOpenAccess":false,"openAccessPdf":"https://onlinelibrary.wiley.com/doi/epdf/10.1002/cpz1.1125","citationCount":"0","resultStr":null,"platform":"Semanticscholar","paperid":null,"PeriodicalName":"Current protocols","FirstCategoryId":"1085","ListUrlMain":"https://onlinelibrary.wiley.com/doi/10.1002/cpz1.1125","RegionNum":0,"RegionCategory":null,"ArticlePicture":[],"TitleCN":null,"AbstractTextCN":null,"PMCID":null,"EPubDate":"","PubModel":"","JCR":"","JCRName":"","Score":null,"Total":0}
引用次数: 0
Abstract
In vitro amplification of single-stranded oligonucleotide libraries presents a significant challenge due to the potential for excessive byproduct formation. This phenomenon largely affects the quality of the ssDNAs created using the most commonly used methods, e.g., asymmetric PCR, biotin-streptavidin separation, or lambda exonuclease digestion of dsDNA. Here, we describe an improved protocol that combines primer-blocked asymmetric PCR (PBA-PCR) with emulsion PCR and a cost-effective downstream process that altogether alleviates byproduct formation without distorting the sequence space of the ssDNA library. In PBA-PCR, the reaction mixture is complemented with a 3′-phosphate-blocked limiting primer that decreases mispriming, thus reducing polymerization of DNA byproducts. The downstream process includes mixing of the PBA-PCR product with excess reverse complement of the 3′-phosphate-blocked limiting primer and removal of dsDNA strands via biotin-streptavidin separation, yielding purified ssDNAs. In conclusion, we have devised a universally applicable approach for simple and cost-effective production of ssDNA libraries and unique ssDNA sequences with on-demand labeling. Our protocol could be beneficial for a variety of uses, such as generating aptamer libraries for SELEX, creating unique molecular identifiers for a wide range of sequencing applications, providing donor DNA for CRISPR-Cas9 systems, developing scaffold nanostructures, and enabling DNA-based data storage. © 2024 The Author(s). Current Protocols published by Wiley Periodicals LLC.
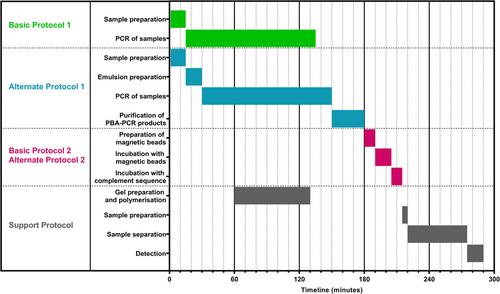
Basic Protocol 1: Amplification of ssDNA libraries using PBA-PCR
Alternate Protocol 1: Amplification of ssDNA libraries using emulsion PBA-PCR with a simplified extraction of PBA-PCR products
Basic Protocol 2: Purification of PBA-PCR products to remove dsDNA and conversion of 3′-blocked primer to double-stranded complexes
Alternate Protocol 2: Purification of PBA-PCR products to remove both dsDNA and blocking primers from the reaction mixture
Support Protocol: Analysis of PBA-PCR products by gel electrophoresis
利用引物阻断非对称 PCR 进行单链 DNA 扩增的经济高效方法
单链寡核苷酸文库的体外扩增是一项重大挑战,因为可能会形成过多的副产物。这种现象在很大程度上影响了使用最常用方法(如非对称 PCR、生物素-链霉亲和素分离或λ外切酶消化 dsDNA)生成的 ssDNA 的质量。在这里,我们介绍了一种改进的方案,它将引物阻断非对称 PCR(PBA-PCR)与乳液 PCR 和一种具有成本效益的下游工艺结合在一起,既能减少副产物的形成,又不会扭曲 ssDNA 文库的序列空间。在 PBA-PCR 中,反应混合物中加入了 3'- 磷酸盐阻断的限制引物,可减少误引物,从而减少 DNA 副产物的聚合。下游流程包括将 PBA-PCR 产物与过量的 3'- 磷酸受限引物反向互补混合,并通过生物素-链霉亲和素分离去除 dsDNA 链,从而得到纯化的 ssDNA。总之,我们设计出了一种普遍适用的方法,可以简单而经济地生产 ssDNA 文库和按需标记的独特 ssDNA 序列。我们的方案可用于多种用途,例如为 SELEX 生成适配体文库、为广泛的测序应用创建独特的分子标识符、为 CRISPR-Cas9 系统提供供体 DNA、开发支架纳米结构以及实现基于 DNA 的数据存储。© 2024 作者。当前协议》由 Wiley Periodicals LLC 出版。基本方案 1:使用 PBA-PCR 扩增 ssDNA 文库 替代方案 1:使用乳液 PBA-PCR 扩增 ssDNA 文库,简化 PBA-PCR 产物的提取 基本方案 2:纯化 PBA-PCR 产物以去除 dsDNA 并将 3'- 阻断引物转化为双链复合物 替代方案 2:纯化 PBA-PCR 产物以去除反应混合物中的 dsDNA 和阻断引物 支持方案:通过凝胶电泳分析 PBA-PCR 产物。
本文章由计算机程序翻译,如有差异,请以英文原文为准。


 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


