Deep learning large-scale drug discovery and repurposing
IF 12
Q1 COMPUTER SCIENCE, INTERDISCIPLINARY APPLICATIONS
引用次数: 0
Abstract
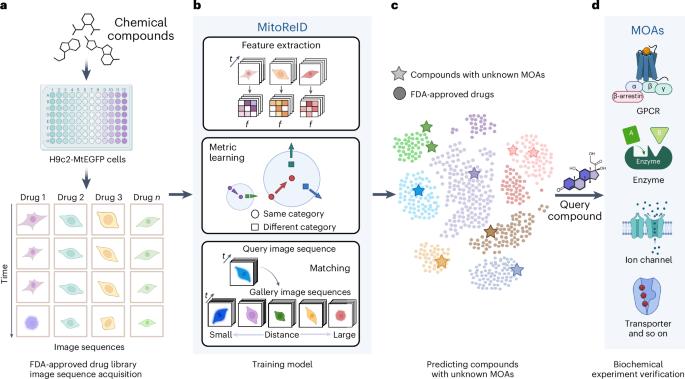
Large-scale drug discovery and repurposing is challenging. Identifying the mechanism of action (MOA) is crucial, yet current approaches are costly and low-throughput. Here we present an approach for MOA identification by profiling changes in mitochondrial phenotypes. By temporally imaging mitochondrial morphology and membrane potential, we established a pipeline for monitoring time-resolved mitochondrial images, resulting in a dataset comprising 570,096 single-cell images of cells exposed to 1,068 United States Food and Drug Administration-approved drugs. A deep learning model named MitoReID, using a re-identification (ReID) framework and an Inflated 3D ResNet backbone, was developed. It achieved 76.32% Rank-1 and 65.92% mean average precision on the testing set and successfully identified the MOAs for six untrained drugs on the basis of mitochondrial phenotype. Furthermore, MitoReID identified cyclooxygenase-2 inhibition as the MOA of the natural compound epicatechin in tea, which was successfully validated in vitro. Our approach thus provides an automated and cost-effective alternative for target identification that could accelerate large-scale drug discovery and repurposing. A deep learning-based model, MitoReID, is presented for profiling changes in mitochondrial phenotypes, allowing for the identification of various drugs’ mechanism of action.
深度学习大规模药物发现和再利用。
大规模药物发现和再利用具有挑战性。确定药物的作用机制(MOA)至关重要,但目前的方法成本高、通量低。在这里,我们提出了一种通过线粒体表型变化进行MOA鉴定的方法。通过对线粒体形态和膜电位进行时间成像,我们建立了一个用于监测时间分辨线粒体图像的管道,形成了一个由 570,096 张单细胞图像组成的数据集,这些图像是暴露于 1,068 种美国食品药品管理局批准药物的细胞的图像。利用重新识别(ReID)框架和膨胀三维 ResNet 主干网,开发了名为 MitoReID 的深度学习模型。该模型在测试集上取得了 76.32% 的 Rank-1 和 65.92% 的平均精度,并根据线粒体表型成功识别了六种未经训练药物的 MOA。此外,MitoReID还确定了茶叶中天然化合物表儿茶素的MOA为环氧化酶-2抑制,并成功地进行了体外验证。因此,我们的方法为靶点识别提供了一种自动化、经济高效的替代方法,可以加速大规模药物发现和再利用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


