Identification of leukemia stem cell subsets with distinct transcriptional, epigenetic and functional properties
IF 12.8
1区 医学
Q1 HEMATOLOGY
引用次数: 0
Abstract
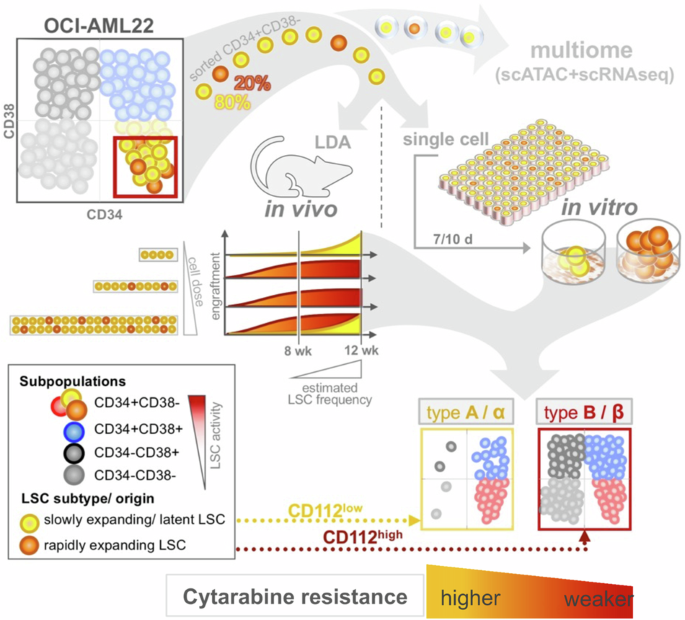
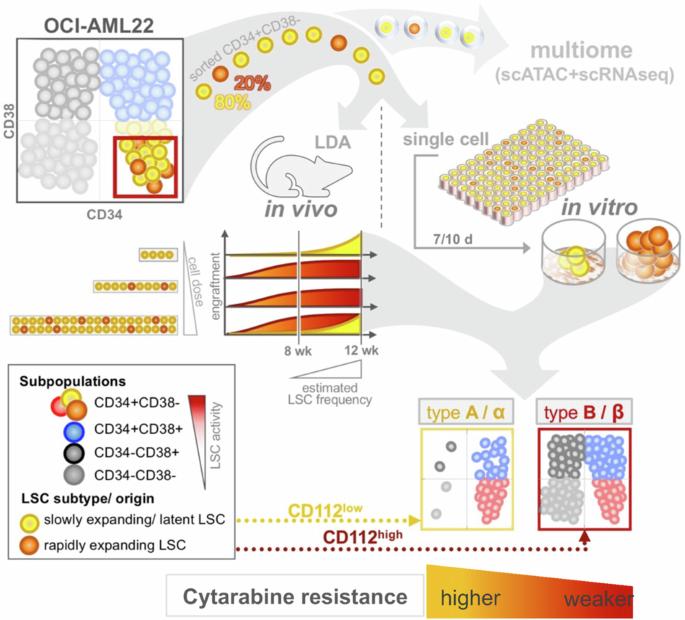
The leukemia stem cell (LSC) compartment is a complex reservoir fueling disease progression in acute myeloid leukemia (AML). The existence of heterogeneity within this compartment is well documented but prior studies have focused on genetic heterogeneity without being able to address functional heterogeneity. Understanding this heterogeneity is critical for the informed design of therapies targeting LSC, but has been hampered by LSC scarcity and the lack of reliable cell surface markers for viable LSC isolation. To overcome these challenges, we turned to the patient-derived OCI-AML22 cell model. This model includes functionally, transcriptionally and epigenetically characterized LSC broadly representative of LSC found in primary AML samples. Focusing on the pool of LSC, we used an integrated approach combining xenograft assays with single-cell analysis to identify two LSC subtypes with distinct transcriptional, epigenetic and functional properties. These LSC subtypes differed in depth of quiescence, differentiation potential, repopulation capacity, sensitivity to chemotherapy and could be isolated based on CD112 expression. A majority of AML patient samples had transcriptional signatures reflective of either LSC subtype, and some even showed coexistence within an individual sample. This work provides a framework for investigating the LSC compartment and designing combinatorial therapeutic strategies in AML.
鉴定具有不同转录、表观遗传和功能特性的白血病干细胞亚群
急性髓性白血病(AML)中,白血病干细胞(LSC)区是一个复杂的储库,对疾病进展起着推波助澜的作用。这一区室中存在的异质性已得到充分证实,但之前的研究主要集中在遗传异质性上,而无法解决功能异质性问题。了解这种异质性对于设计针对造血干细胞的知情疗法至关重要,但造血干细胞的稀缺性和缺乏可靠的细胞表面标志物来分离有活力的造血干细胞一直阻碍着研究的进行。为了克服这些挑战,我们转而使用源自患者的 OCI-AML22 细胞模型。该模型包括具有功能、转录和表观遗传特征的 LSC,广泛代表了原发性 AML 样本中发现的 LSC。我们以LSC池为重点,采用异种移植检测与单细胞分析相结合的综合方法,鉴定出两种具有不同转录、表观遗传和功能特性的LSC亚型。这些LSC亚型在静止深度、分化潜能、再繁殖能力、对化疗的敏感性等方面存在差异,并可根据CD112的表达进行分离。大多数急性髓细胞性白血病患者样本的转录特征反映了其中任何一种 LSC 亚型,有些甚至在单个样本中显示出共存性。这项工作为研究 LSC 区系和设计 AML 组合治疗策略提供了一个框架。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Leukemia
医学-血液学
CiteScore
18.10
自引率
3.50%
发文量
270
审稿时长
3-6 weeks
期刊介绍:
Title: Leukemia
Journal Overview:
Publishes high-quality, peer-reviewed research
Covers all aspects of research and treatment of leukemia and allied diseases
Includes studies of normal hemopoiesis due to comparative relevance
Topics of Interest:
Oncogenes
Growth factors
Stem cells
Leukemia genomics
Cell cycle
Signal transduction
Molecular targets for therapy
And more
Content Types:
Original research articles
Reviews
Letters
Correspondence
Comments elaborating on significant advances and covering topical issues
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


