Genetic and multi-omic risk assessment of Alzheimer's disease implicates core associated biological domains
Abstract
INTRODUCTION
Alzheimer's disease (AD) is the predominant dementia globally, with heterogeneous presentation and penetrance of clinical symptoms, variable presence of mixed pathologies, potential disease subtypes, and numerous associated endophenotypes. Beyond the difficulty of designing treatments that address the core pathological characteristics of the disease, therapeutic development is challenged by the uncertainty of which endophenotypic areas and specific targets implicated by those endophenotypes to prioritize for further translational research. However, publicly funded consortia driving large-scale open science efforts have produced multiple omic analyses that address both disease risk relevance and biological process involvement of genes across the genome.
METHODS
Here we report the development of an informatic pipeline that draws from genetic association studies, predicted variant impact, and linkage with dementia associated phenotypes to create a genetic risk score. This is paired with a multi-omic risk score utilizing extensive sets of both transcriptomic and proteomic studies to identify system-level changes in expression associated with AD. These two elements combined constitute our target risk score that ranks AD risk genome-wide. The ranked genes are organized into endophenotypic space through the development of 19 biological domains associated with AD in the described genetics and genomics studies and accompanying literature. The biological domains are constructed from exhaustive Gene Ontology (GO) term compilations, allowing automated assignment of genes into objectively defined disease-associated biology. This rank-and-organize approach, performed genome-wide, allows the characterization of aggregations of AD risk across biological domains.
RESULTS
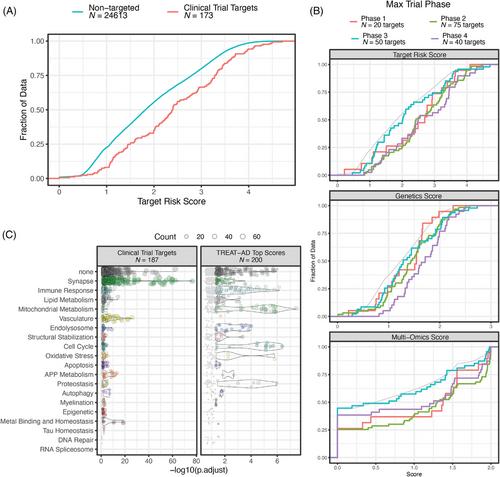
The top AD-risk-associated biological domains are Synapse, Immune Response, Lipid Metabolism, Mitochondrial Metabolism, Structural Stabilization, and Proteostasis, with slightly lower levels of risk enrichment present within the other 13 biological domains.
DISCUSSION
This provides an objective methodology to localize risk within specific biological endophenotypes and drill down into the most significantly associated sets of GO terms and annotated genes for potential therapeutic targets.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


