A 5′ UTR language model for decoding untranslated regions of mRNA and function predictions
IF 23.9
1区 计算机科学
Q1 COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE
引用次数: 0
Abstract
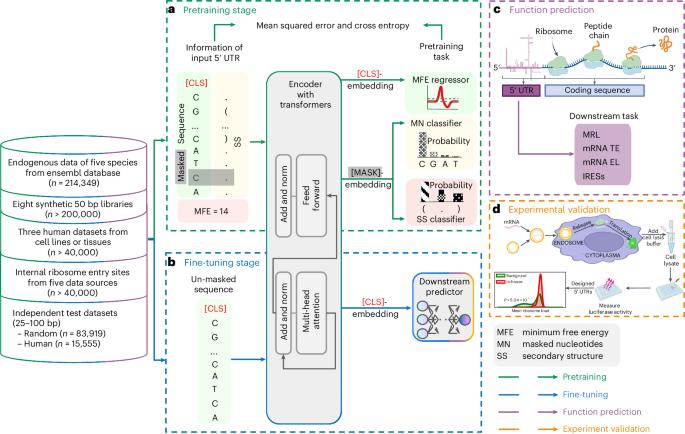
The 5′ untranslated region (UTR), a regulatory region at the beginning of a messenger RNA (mRNA) molecule, plays a crucial role in regulating the translation process and affects the protein expression level. Language models have showcased their effectiveness in decoding the functions of protein and genome sequences. Here, we introduce a language model for 5′ UTR, which we refer to as the UTR-LM. The UTR-LM is pretrained on endogenous 5′ UTRs from multiple species and is further augmented with supervised information including secondary structure and minimum free energy. We fine-tuned the UTR-LM in a variety of downstream tasks. The model outperformed the best known benchmark by up to 5% for predicting the mean ribosome loading, and by up to 8% for predicting the translation efficiency and the mRNA expression level. The model was also applied to identifying unannotated internal ribosome entry sites within the untranslated region and improved the area under the precision–recall curve from 0.37 to 0.52 compared to the best baseline. Further, we designed a library of 211 new 5′ UTRs with high predicted values of translation efficiency and evaluated them via a wet-laboratory assay. Experiment results confirmed that our top designs achieved a 32.5% increase in protein production level relative to well-established 5′ UTRs optimized for therapeutics. The 5′ untranslated region is a critical regulatory region of mRNA, influencing gene expression regulation and translation. Chu, Yu and colleagues develop a language model for analysing untranslated regions of mRNA. The model, pretrained on data from diverse species, enhances the prediction of mRNA translation activities and has implications for new vaccine design.
用于 mRNA 非翻译区解码和功能预测的 5′ UTR 语言模型
5′ 非翻译区(UTR)是信使 RNA(mRNA)分子开头的一个调节区,在调节翻译过程和影响蛋白质表达水平方面起着至关重要的作用。语言模型在解码蛋白质和基因组序列的功能方面展示了其有效性。在这里,我们介绍一种针对 5′ UTR 的语言模型,我们称之为 UTR-LM。UTR-LM 在多个物种的内源 5′ UTR 上进行了预训练,并利用二级结构和最小自由能等监督信息进一步增强。我们在各种下游任务中对 UTR-LM 进行了微调。在预测平均核糖体负载量方面,该模型优于已知最佳基准模型达 5%;在预测翻译效率和 mRNA 表达水平方面,优于已知最佳基准模型达 8%。该模型还被用于识别非翻译区内未注释的内部核糖体进入位点,与最佳基准相比,精确度-召回曲线下的面积从 0.37 提高到了 0.52。此外,我们还设计了一个由 211 个具有较高翻译效率预测值的新 5′ UTR 组成的文库,并通过湿实验室实验对其进行了评估。实验结果证实,与经过优化的成熟 5′ UTR 相比,我们的顶级设计使蛋白质生产水平提高了 32.5%。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Machine Intelligence
Multiple-
CiteScore
36.90
自引率
2.10%
发文量
127
期刊介绍:
Nature Machine Intelligence is a distinguished publication that presents original research and reviews on various topics in machine learning, robotics, and AI. Our focus extends beyond these fields, exploring their profound impact on other scientific disciplines, as well as societal and industrial aspects. We recognize limitless possibilities wherein machine intelligence can augment human capabilities and knowledge in domains like scientific exploration, healthcare, medical diagnostics, and the creation of safe and sustainable cities, transportation, and agriculture. Simultaneously, we acknowledge the emergence of ethical, social, and legal concerns due to the rapid pace of advancements.
To foster interdisciplinary discussions on these far-reaching implications, Nature Machine Intelligence serves as a platform for dialogue facilitated through Comments, News Features, News & Views articles, and Correspondence. Our goal is to encourage a comprehensive examination of these subjects.
Similar to all Nature-branded journals, Nature Machine Intelligence operates under the guidance of a team of skilled editors. We adhere to a fair and rigorous peer-review process, ensuring high standards of copy-editing and production, swift publication, and editorial independence.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


