Oomycete composition in Proteaceae orchards and natural stands on three continents
IF 3
3区 生物学
Q3 MYCOLOGY
引用次数: 0
Abstract
Abstract The Proteaceae , a diverse family of woody flowering plants in the Southern Hemisphere, contains many species known to be susceptible to Phytophthora cinnamomi , both in the natural environment and in cut-flower orchards. Very little is known about the prevalence of P. cinnamomi and other oomycetes across these landscapes. To address this knowledge gap, we used a double ITS1 and RPS10 gene metabarcoding approach and traditional isolation protocols to investigate oomycetes in orchards and natural stands of Proteaceae across South Africa, South Africa (eastern and western), Australia, and Europe. The RPS10 primers amplified more samples, including various Pythium species, while the ITS primers detected more Phytophthora phylotypes. Both datasets showed that geographic regions influenced oomycete species richness and community composition, while they did not show any variation between orchards and natural vegetation. RPS10 metabarcoding detected the largest number of species and provided greater statistical confidence than ITS1 when considering oomycete species composition. Metabarcoding also showed that orchards had a higher abundance of P. cinnamomi compared to native stands, although this was not found when isolating through baiting of roots and rhizosphere soil. Direct isolation and metabarcoding are complementary, with metabarcoding serving as an early detection tool. However, it cannot distinguish living viable propagules from residual DNA of dead propagules, limiting its use for diagnostic purposes related to Phytophthora management and control. These results, along with those of other recent studies, show that metabarcoding offers an effective tool to describe the dynamics of soil oomycetes in different ecosystems.
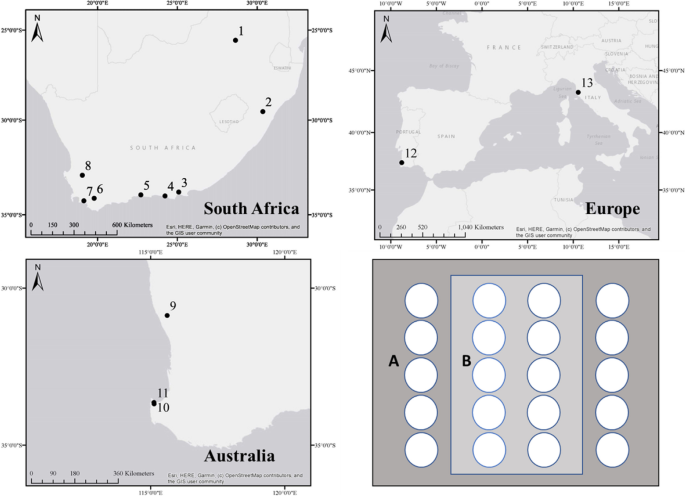
三大洲Proteaceae果园和天然林分的卵菌组成
Proteaceae是南半球木本开花植物的一个多样化家族,包含许多已知在自然环境和切花果园中对Phytophthora cinnamomi敏感的物种。关于肉桂假单胞菌和其他卵菌在这些景观中的流行情况知之甚少。为了解决这一知识空白,我们使用双ITS1和RPS10基因元条形码方法和传统的分离方法研究了南非、南非(东部和西部)、澳大利亚和欧洲的Proteaceae果园和自然林分中的卵菌。RPS10引物扩增的样品较多,包括多种霉属,而ITS引物扩增的疫霉菌种型较多。两组数据均显示地理区域对卵菌种类丰富度和群落组成有影响,而果园与自然植被之间没有差异。RPS10元条形码检测到的物种数量最多,在考虑卵菌种类组成时比ITS1提供了更大的统计置信度。元条形码还显示,与本地林分相比,果园中肉桂的丰度更高,尽管通过根际土壤和根际土壤进行分离时没有发现这一点。直接隔离和元条形码是互补的,元条形码可以作为早期检测工具。然而,它不能区分活的有活力的繁殖体和死亡繁殖体的残余DNA,限制了它在与疫霉菌管理和控制有关的诊断目的中的应用。这些结果以及最近的其他研究表明,元条形码为描述不同生态系统中土壤卵菌的动态提供了有效的工具。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Mycological Progress
生物-真菌学
CiteScore
4.50
自引率
8.30%
发文量
94
审稿时长
6-12 weeks
期刊介绍:
Mycological Progress publishes papers on all aspects of fungi, including lichens. While Review Papers are highly welcome, the main focus is on Research Articles on
Taxonomy and Systematics
Evolution
Cell Biology
Ecology
Biotechnology
Pathology (plants, animals, humans)
Manuscripts on current methods applied in, e.g., morphology, anatomy, ultrastructure (TEM, SEM), genetics, molecular biology, chemistry, and physiology will also be considered.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


