Inconsistent estimates of hybridization frequency in newts revealed by SNPs and microsatellites
IF 1.7
3区 环境科学与生态学
Q2 BIODIVERSITY CONSERVATION
引用次数: 0
Abstract
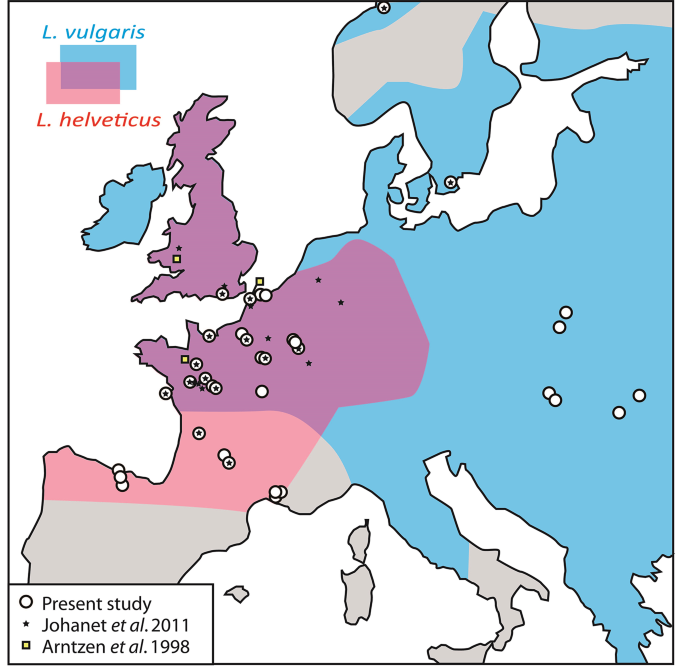
Abstract Hybridization between the European smooth and palmate newts has recurrently been mentioned in the literature. The only two studies that attempted to quantify the frequency of hybridization and gene admixture between these two species came to strikingly opposite conclusions. According to Arntzen et al. (1998, 42 allozymes), hybrids are rare in nature and introgression negligible, while according to Johanet et al. (2011, 6 microsatellites), introgressive hybridization is significant and widespread across the shared distribution range. To clarify this question, we implemented high-throughput SNP genotyping with diagnostic biallelic SNPs on 965 specimens sampled across Europe. Our results are in line with Arntzen et al., since only two F1 hybrids were identified in two distinct French localities, and no further hybrid generations or backcrosses. Moreover, reanalysis of 78 of the samples previously studied by Johanet et al. (2011) using our SNPs panel could not reproduce their results, suggesting that microsatellite-based inference overestimated the hybridization frequency between these two species. Since we did not detect methodological issues with the analyses of Johanet et al., our results suggest that SNP approaches outperform microsatellite-based assessments of hybridization frequency, and that conclusions previously published on this topic with a small number of microsatellite loci should be taken with caution, and ideally be repeated with an increased genomic coverage.
单核苷酸多态性和微卫星显示的蝾螈杂交频率估计不一致
摘要欧洲光滑蝾螈和棕榈蝾螈之间的杂交在文献中经常被提到。仅有的两项试图量化这两个物种之间杂交频率和基因混合的研究得出了截然相反的结论。根据Arntzen等人(1998,42个同工酶)的研究,杂种在自然界中是罕见的,渗入可以忽略不计,而根据Johanet等人(2011,6个微卫星)的研究,在共有的分布范围内,渗入杂交是重要的,并且广泛存在。为了澄清这个问题,我们对欧洲965个样本进行了高通量SNP基因分型,其中包括诊断性双等位基因SNP。我们的结果与Arntzen等人一致,因为在两个不同的法国地区只鉴定了两个F1杂交种,没有进一步的杂交代或回交。此外,使用我们的snp面板对Johanet等人(2011)先前研究的78个样本进行的重新分析无法重现他们的结果,这表明基于微卫星的推断高估了这两个物种之间的杂交频率。由于我们在Johanet等人的分析中没有发现方法学上的问题,我们的结果表明,SNP方法优于基于微卫星的杂交频率评估,并且应该谨慎对待先前发表的关于该主题的少量微卫星位点的结论,最好是在增加基因组覆盖率的情况下重复。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Conservation Genetics
环境科学-生物多样性保护
CiteScore
3.80
自引率
4.50%
发文量
58
审稿时长
1 months
期刊介绍:
Conservation Genetics promotes the conservation of biodiversity by providing a forum for data and ideas, aiding the further development of this area of study. Contributions include work from the disciplines of population genetics, molecular ecology, molecular biology, evolutionary biology, systematics, forensics, and others. The focus is on genetic and evolutionary applications to problems of conservation, reflecting the diversity of concerns relevant to conservation biology. Studies are based on up-to-date technologies, including genomic methodologies. The journal publishes original research papers, short communications, review papers and perspectives.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


