Lipid composition modulates interactions of p7 viroporin during membrane insertion
Abstract
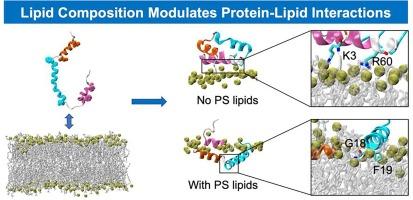
Viral proteins interact with lipid membranes during various stages in the viral life cycle to propagate infection. p7 is an ion channel forming protein of Hepatitis C virus (HCV) that participates in viral assembly. Studies show that it has close ties to lipid metabolism in the cell and anionic phosphatidylserine (PS) lipids are suggested to be key for its permeabilizing function, but the mechanism of its interaction with the lipid environment is largely unknown. To begin unraveling the molecular processes of the protein, we evaluated the impact of lipid environment on the binding and insertion mechanism of p7 prior to channel formation and viral assembly using molecular dynamics simulations. It is seen that p7 is sensitive to its lipid environment and results in different remodeling patterns in membranes. Helix 1 (H1) is especially important for peptide insertion, with deeper entry taking place when the membrane contains phosphatidylserine (PS). Helix 2 (H2) and the adjacent loop connecting to Helix 3 (H3) prompts recruitment of phosphatidylethanolamine (PE) lipids to the protein binding site in membrane models with lower surface charge. This work provides perspectives on the interplay between protein-lipid dynamics and membrane composition, and insights on membrane reorganization in mechanisms of disease.

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


