CNEReg Interprets Ruminant-specific Conserved Non-coding Elements by Developmental Gene Regulatory Network
IF 7.9
2区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
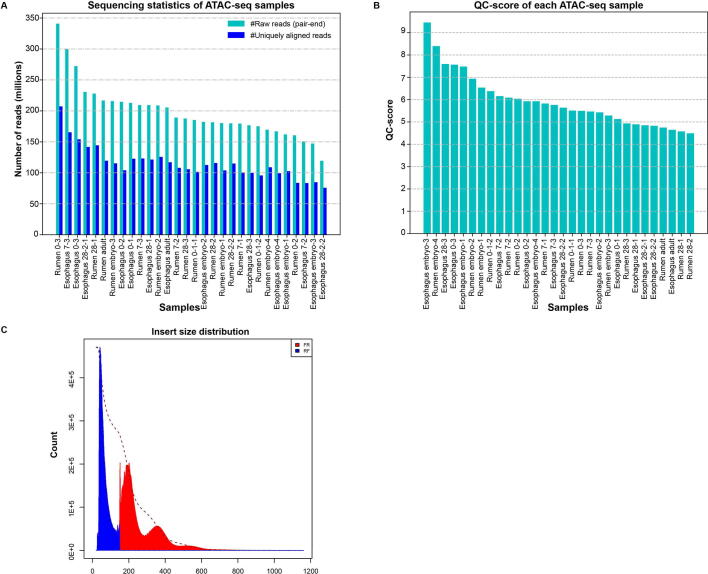
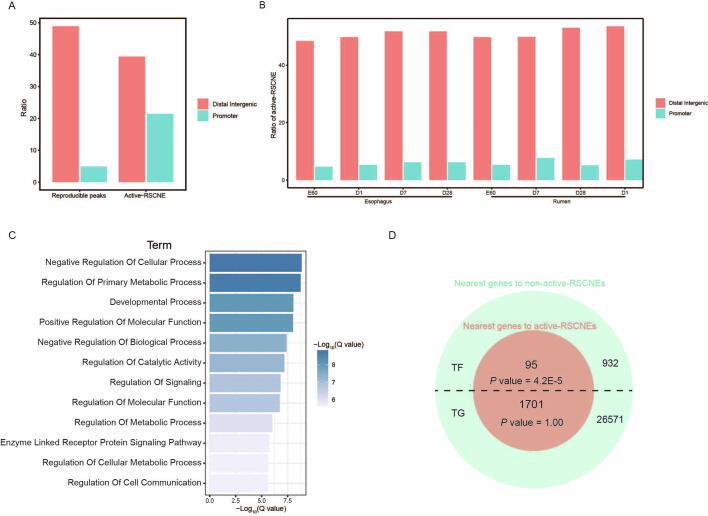
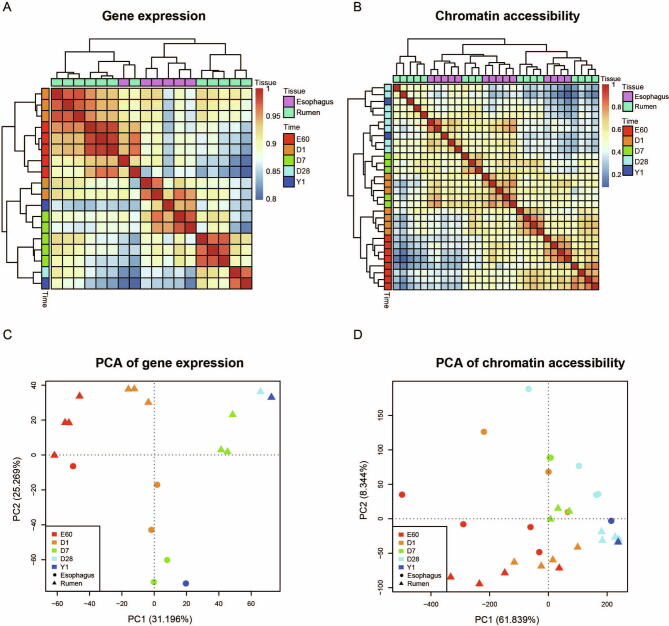
The genetic information coded in DNA leads to trait innovation via a gene regulatory network (GRN) in development. Here, we developed a conserved non-coding element interpretation method to integrate multi-omics data into gene regulatory network (CNEReg) to investigate the ruminant multi-chambered stomach innovation. We generated paired expression and chromatin accessibility data during rumen and esophagus development in sheep, and revealed 1601 active ruminant-specific conserved non-coding elements (active-RSCNEs). To interpret the function of these active-RSCNEs, we defined toolkit transcription factors (TTFs) and modeled their regulation on rumen-specific genes via batteries of active-RSCNEs during development. Our developmental GRN revealed 18 TTFs and 313 active-RSCNEs regulating 7 rumen functional modules. Notably, 6 TTFs (OTX1, SOX21, HOXC8, SOX2, TP63, and PPARG), as well as 16 active-RSCNEs, functionally distinguished the rumen from the esophagus. Our study provides a systematic approach to understanding how gene regulation evolves and shapes complex traits by putting evo-devo concepts into practice with developmental multi-omics data.
CNEReg通过发育基因调控网络解释反刍动物特有的保守非编码元件。
DNA编码的遗传信息在发育过程中通过基因调控网络(GRN)引导性状创新。在此,我们开发了一种保守的非编码元件解释方法,将多组学数据整合到基因调控网络(gene regulatory network, CNEReg)中,以研究反刍动物多室胃创新。我们生成了绵羊瘤胃和食道发育过程中的配对表达和染色质可及性数据,并揭示了1601个活跃的反刍动物特异性保守非编码元件(active- rscnes)。为了解释这些活性- rscnes的功能,我们定义了工具包转录因子(ttf),并在发育过程中通过活性- rscnes细胞模拟了它们对瘤胃特异性基因的调控。我们的发育GRN发现18个ttf和313个活性rscnes调节7个瘤胃功能模块。值得注意的是,6个ttf (OTX1、SOX21、HOXC8、SOX2、TP63和PPARG)以及16个活性rscnes在功能上区分了瘤胃和食管。我们的研究提供了一种系统的方法来理解基因调控是如何进化和塑造复杂性状的,通过将进化-发展概念与发育多组学数据相结合。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Genomics, Proteomics & Bioinformatics
Biochemistry, Genetics and Molecular Biology-Biochemistry
CiteScore
14.30
自引率
4.20%
发文量
844
审稿时长
61 days
期刊介绍:
Genomics, Proteomics and Bioinformatics (GPB) is the official journal of the Beijing Institute of Genomics, Chinese Academy of Sciences / China National Center for Bioinformation and Genetics Society of China. It aims to disseminate new developments in the field of omics and bioinformatics, publish high-quality discoveries quickly, and promote open access and online publication. GPB welcomes submissions in all areas of life science, biology, and biomedicine, with a focus on large data acquisition, analysis, and curation. Manuscripts covering omics and related bioinformatics topics are particularly encouraged. GPB is indexed/abstracted by PubMed/MEDLINE, PubMed Central, Scopus, BIOSIS Previews, Chemical Abstracts, CSCD, among others.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


