解开淡水湖20多年来的病毒生态学和进化
IF 20.5
1区 生物学
Q1 MICROBIOLOGY
引用次数: 0
摘要
随着淡水湖经历快速的人为变化,长期研究揭示了关键的微生物动力学、进化变化和生物地球化学相互作用,但病毒的重要作用仍然被忽视。在这里,利用来自美国威斯康星州门多塔湖的20年时间序列,我们在时间、季节性和环境因素方面表征了130万个病毒基因组。Caudoviricetes类的双链DNA噬菌体在群落中占主导地位。我们鉴定出574个辅助代谢基因家族,代表了超过14万个辅助代谢基因,包括psbA(光合作用)、pmoC(甲烷氧化)和katG(过氧化氢分解)等重要基因,这些基因在几十年和季节中始终存在并活跃。病毒-宿主对(包括keystone Cyanobacteria、methanotrophs和Nanopelagicales)之间的正相关和生态位分化在季节变化中出现。无机碳和铵影响病毒丰度,强调病毒在“自上而下”和“自下而上”相互作用中的作用。进化过程有利于健康基因,减少基因组异质性和优势亚群。这项研究改变了人们对地球微生物群中病毒生态学和进化的理解。本文章由计算机程序翻译,如有差异,请以英文原文为准。
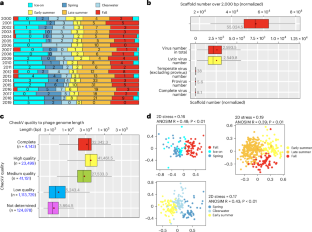

Unravelling viral ecology and evolution over 20 years in a freshwater lake
As freshwater lakes undergo rapid anthropogenic change, long-term studies reveal key microbial dynamics, evolutionary shifts and biogeochemical interactions, yet the vital role of viruses remains overlooked. Here, leveraging a 20 year time series from Lake Mendota, WI, USA, we characterized 1.3 million viral genomes across time, seasonality and environmental factors. Double-stranded DNA phages from the class Caudoviricetes dominated the community. We identified 574 auxiliary metabolic gene families representing over 140,000 auxiliary metabolic genes, including important genes such as psbA (photosynthesis), pmoC (methane oxidation) and katG (hydrogen peroxide decomposition), which were consistently present and active across decades and seasons. Positive associations and niche differentiation between virus–host pairs, including keystone Cyanobacteria, methanotrophs and Nanopelagicales, emerged during seasonal changes. Inorganic carbon and ammonium influenced viral abundances, underscoring viral roles in both ‘top-down’ and ‘bottom-up’ interactions. Evolutionary processes favoured fitness genes, reduced genomic heterogeneity and dominant sub-populations. This study transforms understanding of viral ecology and evolution in Earth’s microbiomes. A long-term metagenomic time series reveals how viruses impact diversity, ecological dynamics and evolution of freshwater microbiomes.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Nature Microbiology
Immunology and Microbiology-Microbiology
CiteScore
44.40
自引率
1.10%
发文量
226
期刊介绍:
Nature Microbiology aims to cover a comprehensive range of topics related to microorganisms. This includes:
Evolution: The journal is interested in exploring the evolutionary aspects of microorganisms. This may include research on their genetic diversity, adaptation, and speciation over time.
Physiology and cell biology: Nature Microbiology seeks to understand the functions and characteristics of microorganisms at the cellular and physiological levels. This may involve studying their metabolism, growth patterns, and cellular processes.
Interactions: The journal focuses on the interactions microorganisms have with each other, as well as their interactions with hosts or the environment. This encompasses investigations into microbial communities, symbiotic relationships, and microbial responses to different environments.
Societal significance: Nature Microbiology recognizes the societal impact of microorganisms and welcomes studies that explore their practical applications. This may include research on microbial diseases, biotechnology, or environmental remediation.
In summary, Nature Microbiology is interested in research related to the evolution, physiology and cell biology of microorganisms, their interactions, and their societal relevance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


